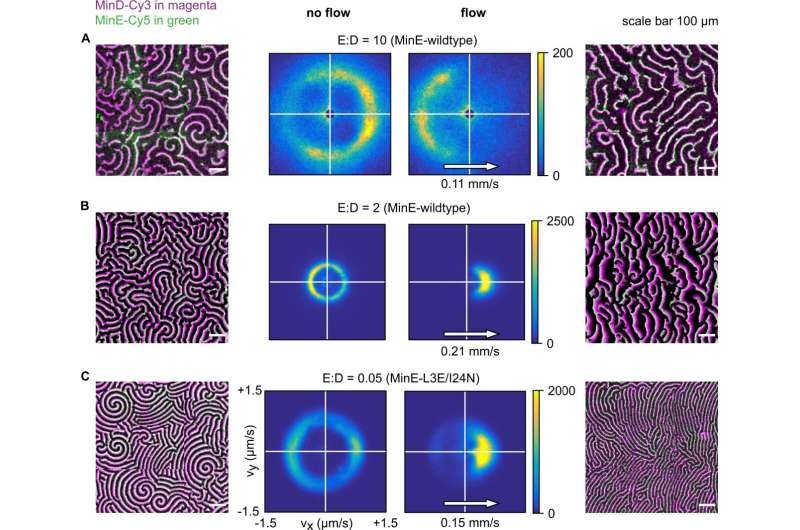
Experimental data showing how patterns respond to flow at different E:D ratios. Data are for MinE-wildtype (A, B) and MinE-L3E/I24N (C), with bulk flow directed left-to-right. Min patterns (outer left and outer right columns) show MinD-Cy3 in magenta and MinE-Cy5 in green. All scale bars are 100 μm. The results of wave-propagation analysis calculated for MinD-Cy3 data are represented as 2D histograms (center columns) with binning size (25 nm/s) × (25 nm/s), showing counts for directionality (vx, vy). Left half of the figures displays an exemplary image as well as wave-propagation analysis for the no-flow case. Right half of the figures displays an exemplary image as well as wave-propagation analysis with flow. Images were stitched from 3 × 3 fields of view. A Upstream propagation was observed in experiments for high E:D ratio (initial 10). B Downstream propagation observed for low E:D ratio (initial 2, corrected 1.3). C Downstream propagation observed for the MinE-L3E/I24N mutant at E:D = 0.05. Credit: Nature Communications (2023). DOI: 10.1038/s41467-023-35997-0
The formation of patterns is a universal phenomenon that underlies fundamental processes in biology. An example are the concentration patterns of proteins, which direct vital cellular processes, including cell division, polarity, and movement. These protein patterns arise from the interplay of chemical reactions and the spatial transport of proteins. Transport can occur either passively (through diffusion) or actively (through flows). Unlike diffusion, transport by flows exhibits a clear preferred spatial direction.
However, little research has been conducted to date into the influence of flows on protein patterns. A team led by LMU physicist Professor Erwin Frey in collaboration with Professor Cees Dekker from the Delft University of Technology has now investigated this fundamental question using the paradigmatic example of the Min protein system of E. coli. The paper is published in the journal Nature Communications.
The researchers used an interdisciplinary approach that combined experiments with microfluidic assays and a novel theory of protein pattern formation developed by the Frey group. In the microfluidic assay the Min proteins can bind and unbind from a lipid bilayer that covers the walls of the microfluidic chamber. This allows researchers to study the behavior of proteins in conditions that closely mimic the inside of an actual biological cell.
In this way, the scientists were able to show that fluid flows cause the movement and alignment of the membrane-bound protein patterns. Surprisingly, the wavelike patterns can move either with the flow direction or against it. Which of these propagation directions occurs depends on the ratio of protein concentrations. The researchers further show how the patterns' direction of movement subtly depends on the chemical reactions between the proteins.
"The membrane-bound patterns can propagate against the flow because the bulk flow shifts the protein concentrations in the cytoplasm but has no direct effects on proteins that are already bound to the membrane," says Frey. "The impact of the flow on protein patterns is determined by how proteins accumulate and deposit on the membrane."
The researchers propose that using fluid flows could be a versatile tool for controlling protein patterns and investigating molecular mechanisms of pattern formation.
More information: Sabrina Meindlhumer et al, Directing Min protein patterns with advective bulk flow, Nature Communications (2023). DOI: 10.1038/s41467-023-35997-0
Journal information: Nature Communications
Provided by Ludwig Maximilian University of Munich