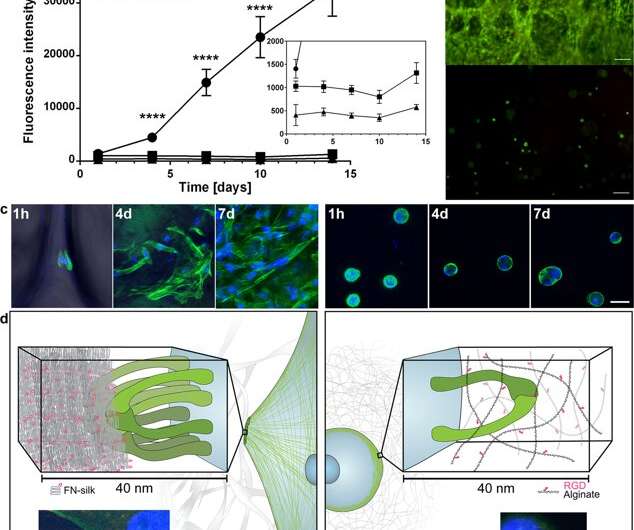
Spreading and expansion of cells within FN-silk compared to when encapsulated in an RGD-coupled hydrogel. (a) Representative graph (mean and standard deviation) of Alamar Blue viability assay showing metabolic activity of fibroblasts (HDF) within FN-silk foam (circle), a very low viscosity (VLVG) alginate hydrogel coupled with RGD (square), and a medium viscosity (MVG) alginate hydrogel coupled with RGD (triangle) during two weeks of culture. Insert shows a zoomed in view of the lower intensities. Statistics (students t-test at each time point): **** indicates p < 0.0001 (n = 12) (b) Representative live (green) and dead (red) staining of human mesenchymal stem cells (HMSC) in FN-silk foam (upper) and RGD-coupled alginate VLVG (lower) at day 14. Scale bars = 100 µm. (c) Confocal scans of HMSCs integrated into FN-silk foam (left panel), and RGD coupled alginate hydrogel MVG (right panel) after 1 h, 4 days and 7 days. Actin filaments are visualized by phalloidin staining (green) and cell nuclei are stained with DAPI (blue). Scale bars = 20 µm. (d) Schematic description of the hypothesis for the reason behind the observed difference in interactions between cells and silk (left) versus RGD-coupled alginate hydrogels (right). Several integrin pairs (green) can adhere and gather to the silk microfibers, forming focal adhesions at the edge of actin filaments, enabling the cells to spread and proliferate. In the alginate hydrogel, a single integrin pair (green) can bind to the coupled RGD-motif, but the thin alginate chains restrict subsequent gathering into focal adhesions. Inserts show examples of a cell (fibroblast) after 3 h in FN-silk foam (left) and a low viscosity (VLVG) alginate hydrogel coupled with RGD (right). Actin filaments are visualized by phalloidin staining (green), and focal adhesions can be seen where this is co-localized with staining for vinculin (red, marked with arrow). Cell nuclei are stained with DAPI (blue). Credit: Scientific Reports, doi: 10.1038/s41598-019-42541-y
Biological tissues are built when cells anchor to specific sites on a 3-D microfiber network in an extracellular matrix (ECM). Scientists are keen to recreate biological tissues in the lab using bioinspired tissue engineering and genetic engineering, to form functional ECM motifs fused to recombinant silk proteins. Under adequate physiological conditions, bioengineered silk proteins and fibronectin-silk (FN-silk) can self-assemble into microfiber networks that mimic native ECM.
In a recent study, Ulrika Johansson, Mona Widhe and co-workers at the interdisciplinary departments of Biotechnology, Biomaterials Chemistry, and Immunology in Sweden developed a method to include mammalian cells into a silk solution before assembling silk into constructs, to form uniform cell-integrated tissue-like microfibers. The resulting 3-D scaffold constructs showed improved cell proliferation (growth) and homogenous cell spreading compared to cells encapsulated in hydrogel. The results of the study are now published in Scientific Reports.
The scientists confirmed cell attachment on fibronectin-silk constructs (FN-Silk) in the work by observing filamentous actin and by defining focal adhesion points of the attached, elongated cells. They maintained cell viability for 90 days in the cell-FN/silk surfaces and showed scalability of the method to macro-sized 3-D cell cultures. The silk microfiber bundles with encapsulated cells maintained biomechanical strength and extendibility much like human arterial walls.
The protocol developed by Johansson and Widhe et al. also allowed stem cells to differentiate inside the 3-D constructs to assist the growth of diverse cell co-cultures. They showed that endothelial cells could be included into the bioinspired materials to form vessel-like structures throughout the tissue constructs. The scientists envision using the ECM-like network as a foundation for future efforts to engineer functional biological tissues in the lab.
In vitro mammalian cell culture is an indispensable experimental technique in basic research and industrial applications, although the existing process relies on 2-D hard plastic or glass surfaces for convenience—impairing the native biological response. Since biological cells are naturally accustomed to receiving signals from the 3-D environment, tissue engineers have formed new experimental strategies using 3-D cell cultures. The experimental conditions maintained cell adhesion, proliferation and differentiation to recreate and sustain cell metabolism and functionality in the lab.
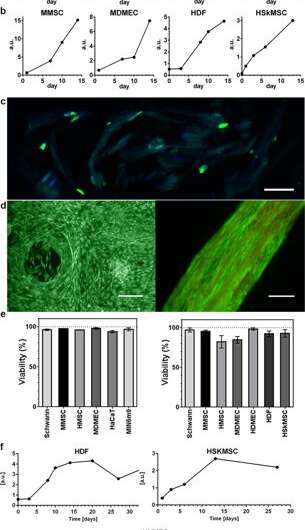
Proliferation and viability of cells integrated in 3D silk. Representative graphs of Alamar blue viability assay show increasing metabolic activity during the first 2 weeks within (a) foam (N = 3–4, n = 3–7), and (b) fibers (N = 1–9, n = 2–13), reflecting growth of the various integrated cell types (HaCaT; human keratinocyte cell line, HDF; human dermal fibroblasts, MDMEC; mouse dermal microvascular endothelial cells, MIN6m9;insulin-secreting mouse pancreatic β-cell line). (c) Cell division occurs deep within the 3D silk. Cryosection of a fiber with integrated fibroblasts (HDF) fixed at day 11 and stained with FITC-anti BrdU for newly synthesized DNA (green) and DAPI (blue). The silk shows a dim autofluorescence in the blue/green range. (d) Representative live (green) and dead (red) staining of mouse mesenchymal stem cells (MMSC) in foam (left) and HDF in fiber (right) at day 14. The fiber shows a dim autofluorescence in the red range. Scale bars = 100 µm. (e) Viability (%, mean and standard deviation) after 14 days culture of different cell types (see Suppl. Table 1) in foam (left graph), and in fibers (right graph) (N = 1–3, n = 4). (f) Long time cultures of cells integrated into fibers maintained metabolic activity (Alamar blue) during the entire study period (up to 97 days). Credit: Scientific Reports, doi: 10.1038/s41598-019-42541-y
Previously, Johansson and Widhe et al. had developed a scalable process to engineer the recombinant spider silk protein known as 4RepCT for bioinspired cell culture in the lab, which self-assembled into biodegradable and biocompatible microfibers in aqueous, physiological buffers at room temperature. They functionalized the novel construct using a cell adhesion motif from fibronectin (FN) to form the FN-silk material and promote firm cell attachment. Although cells proliferated along the new material surfaces, they remained on the surface alone, unable to proliferate in to the constructs to adequately mimic tissue-like properties in vitro. In the present work, the scientists therefore developed a new method, to efficiently embed cells in to the silk material during the assembly of FN-silk for encapsulated and viable 3-D cell culture that adequately mimicked the extracellular matrix in vitro.
Tissue engineering cell-embedded silk constructs in the lab
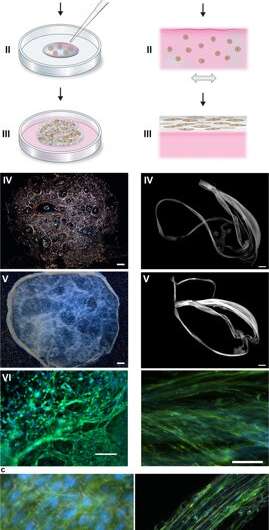
During the experiments, Johansson and Widhe et al., first added a drop of dispersed stem cells (mesenchymal mouse stem cells, MMSC) to the FN-silk protein solution prior to solution assembly. After incubation, the newly formed network remained stable in culture media and the encapsulated cell number increased in the constructs throughout the period of culture. After three days, the cells spread across all dimensions of the foam, which the scientists observed using differential interference contrast (DIC) microscopy.
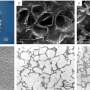
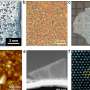
Silk-assembly to integrate cells into a 3D network of microfibers. (a) Schematic description of the formation of silk foam with ingrained cells. Cells suspended in culture medium (pink) are added to a defined drop of FN-silk protein solution (blue) placed in the middle of a non-treated culture well (I). Air bubbles are quickly introduced (5–10 sec) through a pipette tip (II), to give rise to a 3D foam with cells. After 30 minutes in the cell incubator, additional culture medium is added to cover the foam completely (III). Day 1 after formulation, the silk foam with cells looks almost transparent, although harboring some remaining air bubbles (that disappear with time) (IV). After 2 weeks of culture, the foam with integrated cells shows a denser and whiter appearance (V). Already at day 3 the foam is filled with well-spread cells (here mouse mesenchymal stem cells (MMSC) are shown) (VI). Actin filaments are visualized by phalloidin (green) and cell nuclei by DAPI staining (blue). Scale bar IV-V = 1 mm, VI = 100 µm. (b) Schematic description of silk fibers formed with integrated cells. Cells suspended in medium (pink) are added to the FN-silk protein solution (blue) (I). During gentle repeated uniaxial tilting for 1–3 hours (II) the silk proteins assemble at the air-liquid interface into a macroscopic bundle of microfibers with incorporated cells (III). The silk fibers with cells are easily retrieved (IV) and can be placed in a well for further culture, whereby the thickness increases over 2 weeks (V). At day 3, aligned cells (here MMSC) are found spread integrated in the fiber bundle (VI). Actin filaments are visualized by phalloidin (green) and cell nuclei by DAPI staining (blue). Scale bar IV-V = 1 mm, VI = 100 µm. (c) Differential Interference Contrast (DIC) micrographs of the silk microfibers (white) in a fiber (left) and a foam (right) with integrated cells (MMSC). Actin filaments are visualized by phalloidin (green) and cell nuclei by DAPI staining (blue). Scale bar = 50 µm (left) and 100 µm (right). Credit: Scientific Reports, doi: 10.1038/s41598-019-42541-y
They directed the cell alignment to form a random 3-D network of microfibers that successfully mimicked biological tissues by forming a macroscopic bundle of microfibers during the timeline of cell culture, followed by cell to ECM ratio examination. The scientists varied the ECM to cell composition to mimic or match a range of tissue types, including cell sparse and high-density tissues of the liver. The method is therefore also suited for 3-D cell culture with minute cell quantities.
Testing cell viability on the silk scaffolds – cytocompatibility studies
The scientists used growth profiles to map the diverse cell types embedded in both foam and fibers of the silk assembly. They observed an increased signal from the metabolic activity to represent cell proliferation in the 3-D silk scaffolds and in time they showed increased cell density in the innermost cell-silk scaffold. Johansson et al. investigated cell proliferation using BrdU staining, where positive results proved deeper proliferation and cell spreading in the silk fibers to maintain cell viability after 2 weeks of encapsulation, and during long-term cell culture periods that spanned one to three months.
Comparing the cytocompatibility of silk vs. hydrogel biomaterials
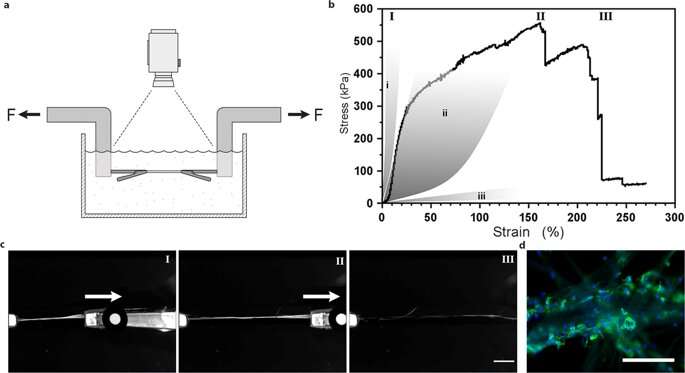
Uni-axial tensile testing of silk fibers with integrated mesenchymal stem cells. (a) Illustration of the experimental set-up for tensile tests performed in PBS buffer at 37 °C in order to maintain viable cells. (b) Representative First Piola-Kirchhoff stress versus strain curve of a FN-silk fiber with integrated mesenchymal stem cells (MMSC) subjected to tensile testing after 14 days of culture. Stress-strain curve illustrates a rather linear (and probably elastic) phase that is followed by a plastic-like (irreversible) deformation phase until the maximum stress is reached, and the fiber breaks. For comparison, grey areas represent ranges of stress strain properties in tendons and ligaments (i), artery walls (ii), and brain tissue (iii). Roman numbers refer to images (c) taken during the tensile test, i.e. during start (I), extension (II) and breakage (III) of the fibers. Scale bar = 5 mm. (d) Micrographs of the breaking point of fibers with MMSCs after tensile testing. Actin filaments are visualized by phalloidin staining (green) and cell nuclei are stained with DAPI (blue). Scale bars = 200 µm. Credit: Scientific Reports, doi: 10.1038/s41598-019-42541-y
Based on the encouraging preliminary results, the scientists conducted parallel experiments to compare cell growth in silk vs. cell growth in hydrogel to determine cytocompatibility of the two materials. They chose alginate to represent the hydrogel during cell culture and observed dissimilarities between alginate vs silk, recording clear cell expansion in silk, while cells in alginate remained in a steady metabolic state. Using confocal microscopy, they investigated reasons for the observed difference in cell growth at the level of the microenvironment. The results showed prompt cell attachment (seen with elongated cells) in the silk constructs, while the alginate constructs contracted during cell culture, which may have stressed the cells to detach.
Biomechanical studies – characterizing the new materials
They determined material surface stiffness to be a crucial component that affected cell fate. To verify this observation, Johansson et al. tested biomechanical behavior of the silk constructs to ensure they adequately mimicked native tissue. They conducted tensile testing in a physiological buffer to obtain the results, which proved that the mechanical properties of silk containing cells matched those of connective tissue such as arterial walls. Johansson et al. were able to demonstrate high extendibility of the microfibers to indicate force transition into and throughout the cells attached to the new biomaterial – confirming adequate cell attachment.
Biofunctionalization studies – investigating surface biocompatibility in vitro
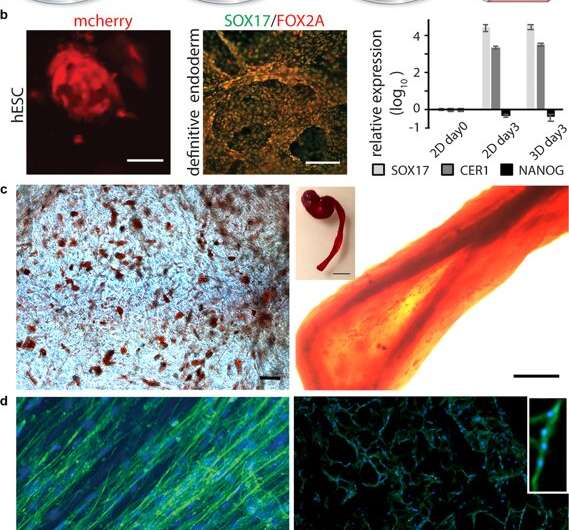
Differentiation of cells in 3D silk. (a) After initial expansion of stem cells integrated to 3D silk, differentiation into various tissue types can be triggered by addition of appropriate factors. (b) Differentiation of pluripotent stem cells. Left: Human embryonic stem cells (hESC) visualized by mCherry detection at 48 h after cell integration into FN-silk foam. Scale bar = 50 µm. Middle: Immunostaining for endodermal markers SOX17 (green) and FOX2A (red) after 3 days of differentiation. Scale bars = 200 µm. Right: Gene expression (SOX17, CER1, NANOG) of hESC in a FN-silk foam compared to 2D culture, analyzed by RT-qPCR at day 3 of endodermal induction. Bars represent the mean fold change ± standard deviation (n = 4). (c) Differentiation of multipotent adult stem cells. Left: Human mesenchymal stem cells (HMSC) in FN-silk foam differentiated into the adipogenic linage containing lipids, visualized by Red Oil staining (red) (N = 2, n = 4). Scale bar = 100 µm. Right: HMSCs differentiated into the osteogenic linage, probed with osteogenic marker for calcium content (Alizarin Red S (red) in FN-silk fiber (right, scale bar = 200 µm), (N = 2, n = 4). Inset shows photo of a whole fiber (right), scale bar = 1 mm). (d) Differentiation of adult precursor cells. Left: After 14 days in differentiation media, skeletal muscle satellite cells (HSkMSC) within a FN-silk fiber show prominent actin filaments, as visualized by phalloidin staining (green). Right: Myogenic differentiation of skeletal muscle satellite cells (HSkMSC) visualized by Desmin staining (green). DAPI-stained nuclei in blue. (N = 9, n = 4). Scale bars = 200 µm. A closeup of the area of a multinucleated myotube is shown in the upper right corner. Credit: Scientific Reports, doi: 10.1038/s41598-019-42541-y
After establishing biomechanical stability, the scientists tested if the stem cells that grew on the silk scaffolds could differentiate (mature) on the same material. For this, they used pluripotent and multipotent human embryonic stem cells in the cell culture assays, followed by quantifying gene expression of biological markers of cell growth; FOXA2 (metabolic genes), SOX17 (genes for embryonic development and cell fate determination), CER1 (bone morphogenetic protein antagonists) and NANOG (embryonic stem cell proliferation, pluripotency and renewal). In the results, both SOX17 and CER1 showed robust upregulation, while the biomarker of pluripotency (NANOG) decreased due to cell maturity as a result of the loss of pluripotency.
The scientists tested surface biofunctionalization for diverse cell types, including human skeletal muscle satellite cells (HSkMSC) and bone marrow-derived human mesenchymal stem cells (hMSC). After cell expansion on the constructs, the scientists could steer the fate of the hMSCs into either adipogenic or osteogenic cell lineages. Additionally, after two weeks of cell culture, Johansson et al. showed myogenic differentiation of the HSkMSCs to form prominent actin filaments, and express the muscle-specific marker desmin, to verify in vitro myotube maturation.
Engineering biological vessels in the lab
The scientists then combined endothelial cells in the 3-D vascular network to form connective tissue that mimicked inherent cellular organization of micro-vessels in the lab. They followed the same protocol using silk assembly with cell integration and added a fraction of endothelial cells to engineer the connective tissue. In two weeks, they observed the cells gather and form millimeter long branched sprouts and vessel-like structures with prominent rings of endothelial cells in the silk fibers. The scientists could increase the size of the constructs to-scale and determine the alignment and aggregation of diverse cell types.
In this way, Johansson and Widhe et al. demonstrated a new strategy and developed a protocol to fit in functional cells within 3-D networks that mimicked the fibrous architecture of the native extracellular matrix (ECM). To mediate the experiment, they used self-assembling recombinant silk proteins and showed that a variety of cells could be embedded in the 3-D constructs. The setup and protocol are simple and cost-effective, unlike 3-D printing the process is frugal and hands-on, without expensive machinery. The scientists aim to optimize and standardize this protocol to develop biocompatible, advanced silk materials in tissue engineering. The experimental work will have a wide range of applications in materials science as miniature in vitro models for drug development and as larger bioengineered tissue constructs in regenerative medicine.
More information: Ulrika Johansson et al. Assembly of functionalized silk together with cells to obtain proliferative 3-D cultures integrated in a network of ECM-like microfibers, Scientific Reports (2019). DOI: 10.1038/s41598-019-42541-y
Brendon M. Baker et al. Cell-mediated fibre recruitment drives extracellular matrix mechanosensing in engineered fibrillar microenvironments, Nature Materials (2015). DOI: 10.1038/nmat4444
Darren Rodenhizer et al. A three-dimensional engineered tumour for spatial snapshot analysis of cell metabolism and phenotype in hypoxic gradients, Nature Materials (2015). DOI: 10.1038/nmat4482
Young's moduli and shear moduli in cortical bone. Proceedings of the Royal Society Series B: Biological Sciences. doi.org/10.1098/rspb.1996.0044
Journal information: Scientific Reports , Nature Materials
Provided by Science X Network
© 2019 Science X Network