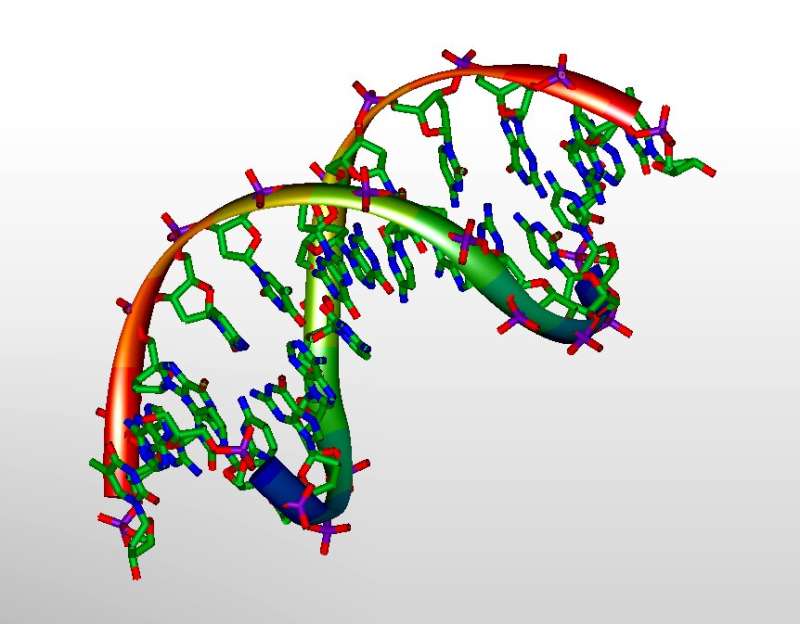
3D-model of DNA. Credit: Michael Ströck/Wikimedia/ GNU Free Documentation License
Nanoscientists and theoretical physicists at UNSW Medicine & Health's EMBL Australia Node in Single Molecule Science joined forces to demystify the complicated mechanisms governing how quickly two matching strands of DNA can fully come together—or hybridize—to form double stranded DNA. Their findings are published in the journal Nucleic Acids Research.
A theory was proposed some 50 years ago hypothesizing that how quickly DNA strands hybridize is determined by the initial contact that leads to further binding of the string of matching bases on the DNA strands—called nucleating interactions. Until now, this theory had never been proven due to the many complexities around DNA biology.
"There are an enormous number of pathways through which two fully dissociated strands can bind to each other. DNA stands don't come together into a fully hybridized duplex in an instant. At some point, only two or three base pairs will spontaneously join. This is what a nucleating event is," said Associate Professor Lawrence Lee who led the team of researchers from UNSW Medicine & Health, UNSW Science, and Imperial College London.
"We built a simple mathematic model, which only has two parameters, and asked: if we only knew how many nucleating interactions there were, and how stable they were, can we predict hybridization rates? And we found that the answer was yes," he said.
To test this model quantitatively, the research team translated the original hypothesis into a mathematical formula that they could use to measure against their experimental observations with synthetic DNA.
A/Prof Lee explains that simplicity was pivotal to the predictive power of their model.
"If a mathematical model contains too many different parameters, it is no longer useful for making predictions. The key difference to previous attempts to understand DNA hybridization rates was that our model had few parameters and was tested against DNA sequences that should not form secondary structures," he said.
DNA secondary structures form when the strands fold onto themselves, which can potentially obscured nucleation and binding sites.
"The theory is, if this initial small interaction is stable enough, it will go from there to a very fast zippering up of the DNA strands. If the limiting step is nucleating, then it follows that if you have more nucleating states, then the DNA should hybridize faster," said A/Prof Lee.
This discovery has the potential to improve our understanding of biological systems. The ability to predict or control the rate of DNA hybridisation, could also help to refine or expand the utility of nanotechnologies. With this new understanding, researchers can adjust the number and stability of nucleation interactions and, in turn, control the rate of DNA binding. This can be achieved in many ways, including by altering the reaction temperature, DNA sequence, and ionic strength of the solution.
"We can generate high resolution images using DNA paint—fluorescent strands of DNA used as tags for microscopy—because we are measuring the binding and unbinding of DNA to individual molecules. But, it can take a long time to acquire data. If we could rationally design sequences for DNA paint, so that it can bind more rapidly, then we could reduce the acquisition time for super-resolution imaging," said A/Prof Lee.
More information: Sophie Hertel et al, The stability and number of nucleating interactions determine DNA hybridization rates in the absence of secondary structure, Nucleic Acids Research (2022). DOI: 10.1093/nar/gkac590
Journal information: Nucleic Acids Research
Provided by University of New South Wales