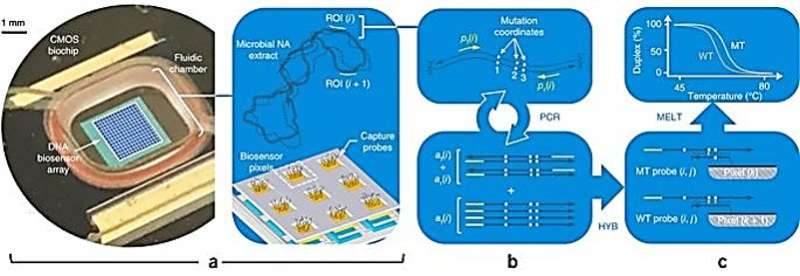
The NAAT platform architecture; a) the closed tube system contains a fluid chamber affixed to a DNA biosensor array, the chamber can receive microbial nucleic acid (NA) extracts and multiplex polymerase chain reaction (PCR) reagents for solution-phase amplification. The DNA capture probes in close affinity to the amplified regions of interest (ROI) in the microbial genome will functionalize individual biosensor pixels in the system, b) each ROI is amplified via PCR to generate amplicons, c) the amplicons bind to specific capture probes on pixels during real-time hybridization (HYB). Thermodynamic stability of these associations is tested with melting curve analysis (MCA) to differentiate between the wild-type and mutant strain signature. Credit: Nature Biotechnology, doi:10.1038/nbt.4179
Evolving strains of multi-drug resistant pathogens are a growing global concern, outpacing drug discovery efforts and undermining the efficacy of existing antibiotics. The development of comprehensive diagnostics for clinical applications will become crucial to control escalating health risks. Existing laboratory tests to diagnose infectious disease are generally carried out via culture-based methods that usually take days to generate results. Rapid molecular diagnostic tests can comparatively identify microbial nucleic acids (NA) in clinical samples directly in less than an hour with nucleic acid amplification tests (NAATs). Existing NAATs are, however, limited by inadequate levels of multiplexing (i.e. the number of strains or sequences detected in a single reaction) and inaccuracies with detecting mutations.
Scientists and engineers recently collaborated to develop a fully integrated, miniaturized semiconductor biochip with closed-tube detection chemistry to perform multiplexed NAAT with aims to overcome existing limitations. The system can rapidly amplify DNA and RNA sequences in a single sample, including species signatures, genotypes and mutations that confer pathogenic drug resistance. Feasibility of the system was first demonstrated by identifying and quantifying multiple viral RNAs and DNAs from respiratory tract pathogens. Subsequently, the system was used to detect > 50 antibiotic resistant mutations across multiple genes of the Mycobacterium tuberculosis (MTB) genome. The research is published in Nature Biotechnology.
In the study, Hassibi et al. designed and developed a miniaturized and single-use (disposable) semiconductor, using conventional sub-micron, complementary metal oxide semiconductor processors (CMOS). The architecture of the biochip contains a fluid chamber with thermocycling capabilities and DNA capture probes with fluorophores that sit above an embedded biosensor array. The fluid chamber of the system can receive multiplex PCR reagents and microbial nucleic acid of a clinical pathogen – with a specific region of interest responsible for drug resistance—for its amplification. When the amplified region of the microbial genome is in close affinity to the DNA capture probe on the biochip, individual biosensor pixels will be activated to detect and differentiate between the wild type and mutant microbial strains.
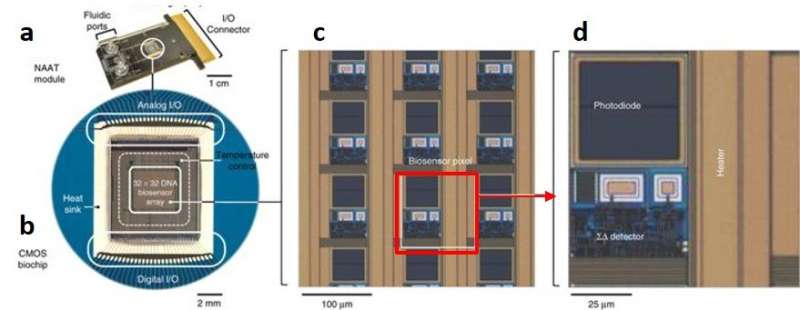
In the NAAT module, fluidic ports can introduce the multiplex PCR mix and extracted nucleic acids into the chamber for nucleic acid amplification. The biochip houses an array of 32 x 32 biosensing pixels, each with embedded photodiode, heaters and detectors. The system is built to identify multiple and distinct nucleic acid sequences in real-time via the inverse fluorescence transduction (IFT) method. By function, the CMOS biochip can detect real-time IFT signals while controlling the temperature of the fluid chamber to enable nucleic acid amplification.
Structure of the biochip and inbuilt components: a) the disposable NAAT module includes a fluidic cap with inlet and outlet ports to b) encapsulate the CMOS biochip mounted on the printed circuit board, c) the biosensor pixels (red highlight) are arranged in a 32 x 32 (1024) array, d) each biosensing pixel is housed with a photodiode, heaters and detectors. Credit: Nature Biotechnology doi: doi:10.1038/nbt.4179
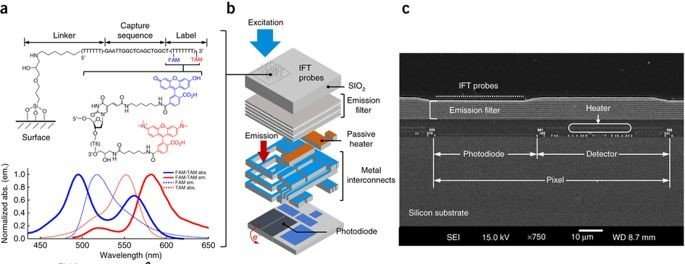
Instead of conventional PCR dyes, the system includes real-time transduction with oligonucleotide IFT probes affixed to each pixel, in addition to the individual photodetector and electronic readout circuits housed within to acquire and digitize fluorescent emissions. Each probe constitutes a dual-fluorophore label to hybridize with PCR amplicon analytes. Overall, the sensor array has an adjustable speed of one to 50 reads per second.
Structure of the biosensor pixels: a) the IFT probes contain a modular linker and a dual-fluorophore label (FAM-TAM) to recognize the PCR amplicon analytes, b) schematic illustration of the 3D stack of the biosensor pixel with localized IFT probes, and b) a scanning electron microscope derived cross-section with emission filters, heater, photodiode and detector of a single pixel fabricated on silicon. Credit: Nature Biotechnology doi: doi:10.1038/nbt.4179
The platform can detect hundreds of different sequences, only limited by the number of pixels on the array. To demonstrate multiplexing capacity, researchers used the CMOS biochip to simultaneously identify multiple DNA and RNA respiratory viruses in the same clinical sample. The platform then identified 54 single nucleotide polymorphism mutations (SNPs), located in six genes to detect drug-resistance-conferring mutations, separating the wild type strains (pathogens sensitive to drugs) from the drug-resistant strains.
Although the system has advantages beyond the existing NAAT platforms to include cost-effective multiplexing, workflow simplicity and <2 hours of time-to-result analysis, the system is hypothesis-driven to only detect currently known mutations. The system does not identify new microbes or new mutations—optimally suited for next-generation sequencing efforts. The existing ability to rapidly detect multiple-drug resistant pathogens and mutations can help address emerging antibiotic resistance by providing actionable data to healthcare providers, on time, for more precisely selected medication.
More information: Multiplexed identification, quantification and genotyping of infectious agents using a semiconductor biochip www.nature.com/articles/Nbt.4179 Hassibi et al, 16 July 2018, Nature Biotechnology
Evaluation of Oxford Nanopore's Minion Sequencing Device for Microbial Whole Genome Sequencing Applications www.nature.com/articles/s41598-018-29334-5 Tyler et al, 19 July 2018
World Health Organization Department of Communicable Disease Surveillance and Response. WHO Global Strategy for Containment of Antimicrobial Resistance (Health Organization, Geneva, 2001) www.who.int/drugresistance/WHO … Strategy_English.pdf
World Health Organization: Global antimicrobial resistance surveillance system (GLASS) report, Early implementation 2016-2017 www.who.int/glass/resources/pu … mentation-report/en/
Better tests, better care: improved diagnostics for infectious diseases www.ncbi.nlm.nih.gov/pubmed/24200831 Caliendo et al, 2013, Oxford Academic, Clinical Infectious Diseases.
Molecular mechanisms of antibiotic resistance www.ncbi.nlm.nih.gov/pubmed/25435309 Blair et al, 2015, Nature Reviews Microbiology.
A CMOS Electromechanical Impedance Spectroscopy (EIS) Biosensor Array ieeexplore.ieee.org/document/5625928/ Manickam et al, 2010, IEEE Transactions on Biomedical Circuits and Systems.
Comparative advantages of mechanical biosensors www.ncbi.nlm.nih.gov/pmc/articles/PMC3839312/ Arlett et al, 2013, Nature Nanotechnology.
A Fully Integrated CMOS Fluorescence Biochip for DNA and RNA testing ieeexplore.ieee.org/document/8082158/ Manickam et al, 2017, IEEE Journal of Solid-State Circuits.
Preparation of electrode-immobilized, redox-modified oligonucleotides for electrochemical DNA and aptamer-based sensing www.ncbi.nlm.nih.gov/pubmed/18007622 Xiao et al, 2007, Nature Protocol.
Journal information: Nature Biotechnology , Clinical Infectious Diseases , Nature Reviews Microbiology , Nature Nanotechnology
© 2018 Phys.org