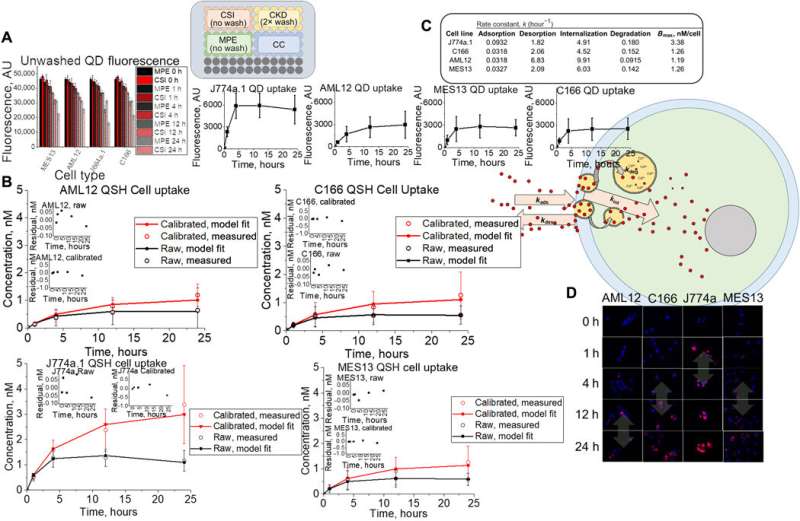
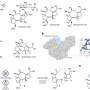
In vitro application of assay using multiple cell types with simulation and rate extraction. (A) Fluorescence data and assay setup representing unwashed QD (QSH) fluorescence for unwashed wells that contain kidney (MES13), liver (AML12), endothelial (C166), and macrophage (J774a.1) cell types. AU, arbitrary units. (B) Washed raw (black) and calibrated (red) fluorescence uptake concentrations (circles, measured) compared to simulation fits (lines) for liver, endothelial, macrophage, and kidney cell types. (C) Summary of rate constants optimized by the genetic algorithm for model fit to measured datasets. (D) Time-dependent confocal microscopy images of QD uptake, showing saturation (double-sided arrows) for each cell type in the study (blue and red represent cell nucleus Hoechst and QSH fluorescence, respectively). Cartoon is a visual representation of QD-cell interaction kinetics. Credit: Science Advances, doi: 10.1126/sciadv.aax2642
Bioengineers can design smart drugs for antibody and nanomaterial-based therapies to optimize drug efficiency for increasingly efficient, early-stage preclinical trials. The ideal drug will have maximum efficiency at target tissue sites for transport from the tissue vasculature to the cellular environment. Researchers can use biological simulations coupled to in vitro approaches to predict their exposure rapidly and efficiently to predict drug biodistribution within single cells of live animal tissue without relying on animal studies. In a new study now published on Science Advances, Edward Price and Andre J. Gesquiere successfully used an in vitro assay and computational fluid dynamic (CFD) model to translate in vitro cell kinetics to whole-body simulations across multiple species and nanomaterial types. The work allowed them to predict drug distributions inside individual tissue cells and the team expect this work to refine, reduce and replace animal testing while providing scientists a fresh perspective on drug development.
Nanomedicines (NMs) in the form of antibodies and synthetic nanomaterials can complement conventional small-molecule medicine through active tissue targeting, variable circulation timeframes and stability, coupled to adjustable biodistribution. Research teams heavily rely on animal models to quantify delivered doses, raising questions on ethics and surges in time and cost. From a scientific viewpoint, tissue architecture destruction conducted during tissue homogenization for quantification can eliminate critical knowledge of nanomedicine transport inside tissue cells and vasculature. A drug can reach cells by exhibiting optimal plasma pharmacokinetics to reach the tissue vasculature. The architecture and nanoparticle diameter allow critical transport of the drug across microvessel walls into an infected or normal tissue cell environment.
In the present work, Price and Gesquiere coupled an in vitro/in silico simulation approach to predict NM biodistribution within preclinical species (animal and cell models) at the level of the single cell. They accounted differences in nanoparticle size, animal species and vasculature pore size diameters using a fluid dynamic model. The team quantified cellular NM content in situations where results were difficult to translate to animal studies due to a lack of systematic perspectives. They also addressed existing experimental issues where cells induced degradation of nanoparticle fluorescence to produce false negatives.
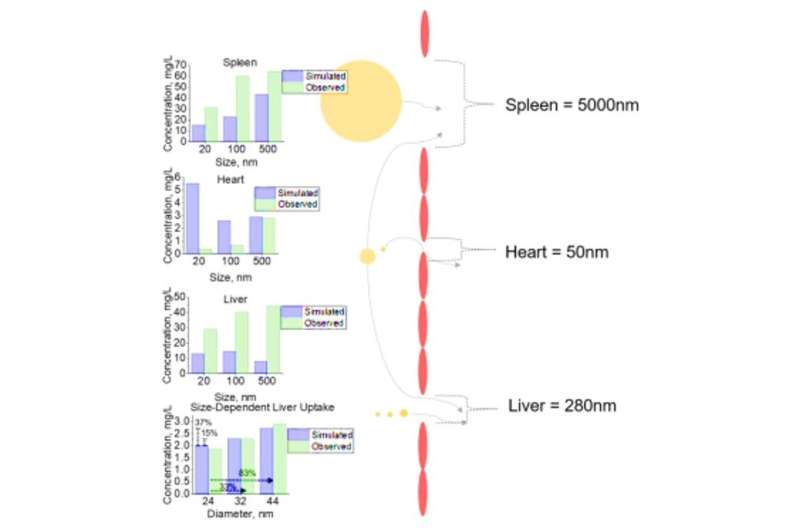
Cellular analysis of NM uptake in varying tissues for NM with different diameters. Cellular uptake of NMs with diameters of 20, 100, and 500nm for spleen, heart, and liver with fenestrae diameters of 5000, 50, and 280nm respectively. Sensitivity analysis of incremental changes in NM diameter (24, 32, and 44nm) with subsequent liver cell uptake were performed. All simulated (predicted) outputs are compared to observed data from literature cellular datasets. Credit: Science Advances, doi: 10.1126/sciadv.aax2642
To validate their work in vitro, the team simulated lysosomal analysis coupled to atomic absorption spectroscopy (AAS), and substantiated in vivo simulations by comparing results to the published literature of whole-body animal data for rats, mice and nonhuman primates. The scientists used quantum dots (QDs) as a model system due to their potential in nanoparticle detection, while also considering their limits. Price et al. accounted the complexities in an NM platform to extend the capabilities to antibody- and metal/polymer-based nanoparticles.
When a nanoparticle circulates through animal blood supply to enter the tissue environment, it will interact with tissue cells via adsorption, desorption, internalization or active uptake processes. The scientists exposed nanoparticles to non-toxic QDs (quantum dots) to fluorescently detect their interactions with cells commonly encountered by NMs such as macrophages, endothelial cells and epithelial cells, after intravenous injection. The research team quantified the concentration of nanoparticles interacting with cells in vitro and built cell kinetics simulations to assess the rate kinetics and NM-cell interactions.
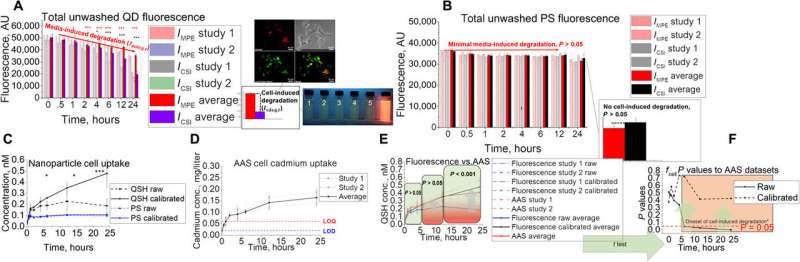
In vitro assay development and validation to AAS. (A and B) In vitro assay setup with its coupled unwashed fluorescence signal for (A) QD and (B) PS. Images show Hepa1-6 colocalization of QD in lysosomal compartments as well as snapshots of fluorescence under ultraviolet illumination under simulated lysosomal exposure conditions with pH 2.5, 3.0, 3.5, 4.0, 4.5, and 7.4 (1 to 6, respectively). (C to E) QD uptake studies using the in vitro technique with (C) fluorescence, (D) AAS, (E) combined, and the (F) statistical results using two-tailed t test when comparing raw and calibrated fluorescence uptake to AAS data. The asterisks in figure represent significance at the * (P < 0.05), ** (P < 0.01), and *** (P < 0.001) levels. Credit: Science Advances, doi: 10.1126/sciadv.aax2642
The simulation contained a (i) medium, (ii) cell membrane and (iii) cell space compartments interconnected via basic mass transfer equations and first-order rate constants. They optimized the system using a genetic algorithm in MATLAB and noted the membrane adsorption rate constant to be highest for macrophages and lowest for endothelial and liver cells. Of the cell types, the internalization process was also relatively highest for liver tissue cells and lowest for endothelial cells. Confocal imaging visually supported these data with rapid uptake and saturation of macrophages within an hour of exposure, while epithelial and endothelial tissue cells were further delayed.
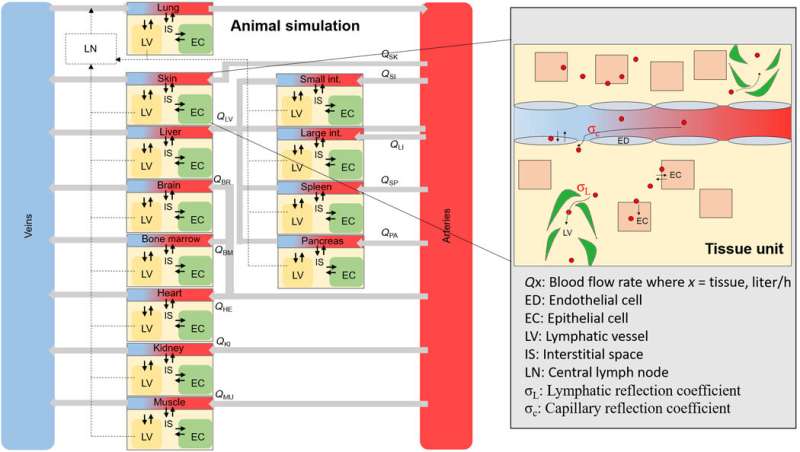
The team demonstrated the presence of cell degradation factors within in vitro assays using cell kinetics simulations to help efficiently translate the in vitro data to animal studies. Atomic adsorption spectroscopy (AAS) validated the results of QD uptake via fluorescence assays and highlighted the importance of calculating cell- and medium-induced degradation. Since nanomedicines (NMs) located inside tissues can accumulate in the interstitia, vasculature or inside a variety of cell types, the team translated in vitro cellular kinetics to in silico animal simulations to account for this. They conducted two pilot studies and assessed trends in whole-tissue uptake as a function of size and quantified NM uptake at the level of the single cell. Architecturally, each tissue compartment simulation contained four sub-compartments to represent the epithelial, endothelial and macrophage cells.
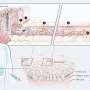
An overall schematic of the animal NM simulation. Scheme of the overall tissue and blood compartments that make up the full animal. Each tissue compartment is interconnected through blood flow rates and contains sub-compartments (zoomed-in area), where an NM will distribute upon internalization through endothelial fenestrae. Credit: Science Advances, doi: 10.1126/sciadv.aax2642
Since the liver and spleen are common targets for NM accumulation, they used these tissues as case studies to understand and capture, the sensitivity and accuracy of the simulation. For example, the simulation showed that increased NM sizes exceeded the tissue pore sizes of most tissue types to funnel toward tissues with larger pore sizes such as the liver. At this point, the liver endothelial cells interacted with the NMs for rapid membrane binding and saturation within 24 hours. When the NM entered the interstitia of the tissue it quickly interacted with epithelial (for liver) and macrophage (for spleen) tissue cells. The results built confidence in the predictive power of the simulation and its capacity to predict variable-sized drug content at the single-cell level for animals for the first time—strictly based on in vitro data alone.
The scientists then indicated the predictive power and translational capacity of the simulation by comparing against measured tissue-level content for multiple NM types and species, including rats, mice and cellular content (epithelial, endothelial and macrophage). They tested the predictive performance across multiple variables based on the World Health Organization guidelines and standards of academic and pharmaceutical consortium for drug development and safety. The model-predicted averages and the observed averages correlated linearly to indicate reasonable model predictions across all species and NM types.
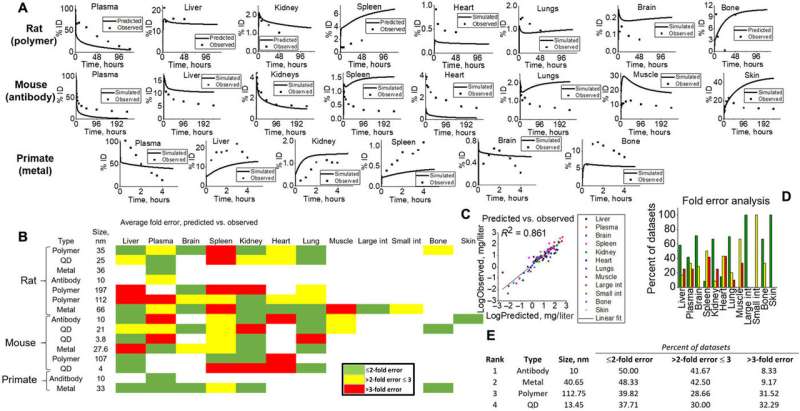
Simulation outputs for validation to various animal studies. (A) Tissue-level predicted (line) versus observed (points) examples for all species types (rats, mice, and NHP) for visual evidence of model capabilities. (B) Heat map of fold error analysis calculated for all tissue, NM, and species types in simulation. Green, yellow, and red represent <2-fold, <3-fold, and >3-fold errors. Fold error was calculated according to equations given in Materials and Methods, where simulated dataset averages were compared to observed. (C) All data point averages specific to tissue types for simulations and observed were plotted against each other to yield a linear fit (R2 = 0.861). (D) Tissue-specific fold errors are shown to visualize model accuracy for each tissue in this study. (E) NM-simulated outputs were ranked (1 to 4) according to fold error analysis. Credit: Science Advances, doi: 10.1126/sciadv.aax2642
In this way, Edward Price and Andre J. Gesquiere developed a viable platform to reduce and refine animal testing during nanomedicine (NM) development. The in vitro concentrated data coupled to cell kinetics simulations delivered cell-NM interactions to show excellent translational potential. The results did not require fitting to animal biodistribution datasets, since all parameters were optimized to in vitro data or calculated using a fluid dynamic model. The process also worked under conditions of NM degradation in the biological environment. The combined in vitro and in silico techniques will assist future smart drug design to help scientists form better and informed discussions while reducing live animal testing. Price and Gesquiere expect this work to serve a new approach for predictive simulations of nanomedicine transport.
More information: Edward Price et al. Animal simulations facilitate smart drug design through prediction of nanomaterial transport to individual tissue cells, Science Advances (2020). DOI: 10.1126/sciadv.aax2642
Matthew Faria et al. Minimum information reporting in bio–nano experimental literature, Nature Nanotechnology (2018). DOI: 10.1038/s41565-018-0246-4
Zheng-Jiang Zhu et al. Stability of quantum dots in live cells, Nature Chemistry (2011). DOI: 10.1038/nchem.1177
Journal information: Science Advances , Nature Nanotechnology , Nature Chemistry
© 2020 Science X Network