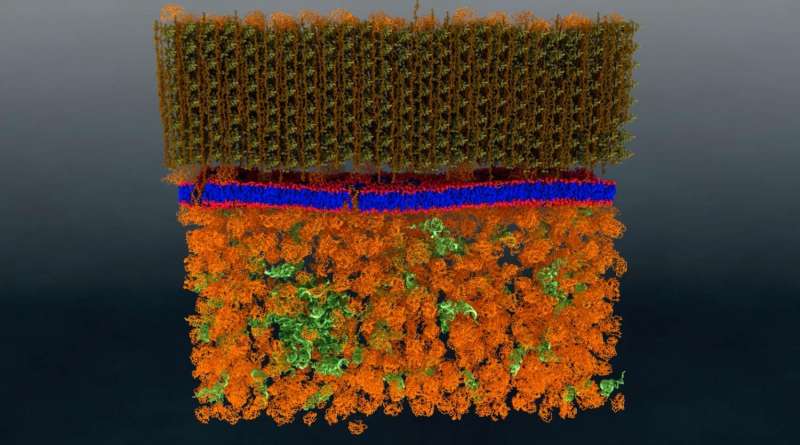
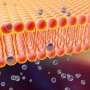
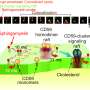
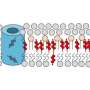
Neutron scattering is a valuable technique for studying cell membranes, but signals from the cell's other components such as proteins, RNA, DNA and carbohydrates can get in the way. An ORNL team made these other components practically invisible to neutrons by combining specific levels of heavy hydrogen (deuterium) with normal hydrogen within the cell. Credit: Oak Ridge National Laboratory
A research team from the Department of Energy's Oak Ridge National Laboratory has performed the first-ever direct nanoscale examination of a living cell membrane. In doing so, it also resolved a long-standing debate by identifying tiny groupings of lipid molecules that are likely key to the cell's functioning.
The methods developed provide a new experimental platform for biophysical studies of membranes and, potentially, other cell components. It could prove useful for future research on important interactions such as drug-membrane, biofuel-membrane, and even antibiotic-membrane interactions.
The multidisciplinary project—led by biophysicist John Katsaras, chemist Bob Standaert and microbiologist James Elkins—was performed at the lab's High Flux Isotope Reactor and Spallation Neutron Source using the bacterium Bacillus subtilis. The team published its findings in the journal PLoS Biology.
A cell's membrane is a thin bilayer of lipid molecules among which reside other biomolecules such as proteins. Researchers have been uncertain about whether membrane lipids sometimes organize into groups called domains, also known as "rafts," or if they are randomly distributed in the membrane. Organization of lipids in distinct domains within the cell membrane is thought to enable functions such as signaling between cells.
"It became a debate," Katsaras said. "Some people believed they exist, while others believed they didn't. There was a lot of circumstantial evidence that could support either side."
The problem was that existing techniques were not capable of unequivocally resolving this question.
Neutron scattering analysis was key to the project's success. Lipid domains are too small to be seen by optical microscopes that use light to probe samples such as biological cells. However, neutrons have no such limitation and can be used to provide a nanoscale view of a cell. Moreover, unlike other nanoscale tools, neutrons can be used for examining a live cell without damaging it.
While neutron scattering analysis overcame the limitations of the other technologies, it presented some daunting challenges of its own. The first was to develop an experiment in which neutrons scattered off of lipid molecules in the membrane without interacting with other components of the cell, such as proteins, RNA, DNA and carbohydrates. The next challenge was to distinguish one type of lipid molecule from another.
The solution to both these challenges lay in the use of deuterium, an isotope of hydrogen whose nucleus contains a neutron as well as a proton. By contrast, common hydrogen nuclei contain a proton but no neutron. While a biological cell itself perceives little difference between normal hydrogen and deuterium, the two isotopes appear very different when looked at using neutron scattering.
The ORNL team created a strain of the bacterium containing enough deuterium to make the cell structures essentially invisible to neutrons. They then ensured that the lipid molecules within the membrane were made up entirely of two fatty acids containing specific proportions of deuterium and hydrogen.
They subsequently introduced the two fatty acids types with differing isotope ratios. The cell membrane was free to create and incorporate into its membrane lipid molecules from them, with each lipid type then containing a specific mix of the two isotopes. If the lipids were distributed randomly throughout the membrane, then the membrane would appear uniform when exposed to neutrons, akin to an optical background that was medium gray.
If, however, the lipids gathered with others of their type, the background would cease to be uniform and would show the equivalent of lighter and darker gray areas. This is in fact what the team found. The grey patches detected using neutrons measured less than 40 nanometers across. The membrane itself was about 2.4 nanometers thick.
The ORNL researchers stressed that their approach of creating internal contrast within living cells using isotopes held promise for other research as well, opening the technique of targeted deuteration to other physical techniques (e.g., nuclear magnetic resonance spectroscopy).
"The people who study these things tend to use particular types of probes," Katsaras noted. "They didn't use neutron scattering because it wasn't in the biologist's wheelhouse. Our novel experimental approach opens up new areas of research.
"For example, you could use the modified bacteria as a platform for investigating antibiotics, because a lot of these antibiotics really are talking to the membrane."
More information: Jonathan D. Nickels et al, The in vivo structure of biological membranes and evidence for lipid domains, PLOS Biology (2017). DOI: 10.1371/journal.pbio.2002214
Journal information: PLoS Biology
Provided by Oak Ridge National Laboratory