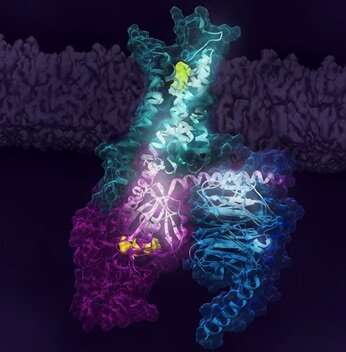
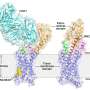
Activation of the human A2A receptor (shown in turquoise) in complex with a G protein consisting of the α, β, and γ subunits (shown in violet, light blue and bark blue, respectively), is exquisitely controlled by changes in dynamics and action of the ligand, highlighted in lime at the top of the figure. Activation pathways are observed to extend from the ligand binding site (lime) to the nucleotide binding site (yellow) in the G protein. Credit: Cell
A multinational research team led by Dr. Adnan Sljoka (RIKEN), Prof. R. Scott Prosser (Univ. of Toronto) with collaborations with Dr. Duy Phuoc Tran and Prof. Akio Kitao (Tokyo Tech) and Prof. Roger K. Sunahara (Univ. of California San Diego) has carried out experimental and computational studies, revealing key steps associated with the activation of the human adenosine A2A receptor (A2AR). A2AR is a member of superfamily of receptors called G protein-coupled receptors (GPCRs; major drug targets) which engage the G protein and initiates cell signaling. The research team discovered that A2AR is represented by at least two inactive conformations and three active conformations whose populations are dependent on ligands and activation states of G protein, and that communication between receptor and G-protein is important for activation and signaling. This study is expected to allow researchers to reach new level of insight in GPCR activation and disease mechanism.
GPCRs affect almost every aspect of human physiology, where 35% of all approved drugs act on GPCRs. In most cases, GPCRs are situated in the plasma membrane that surrounds the cell while the drug or ligand (such as hormones and neurotransmitters) that acts on the GPCR binds to an extracellular pocket. Activation is then transduced across the receptor, resulting in complexation with proteins on the cell interior. Since input arrives at the cell exterior and initiates signaling pathways inside the cell, this makes GPCRs useful in pharmacology as the drug in many cases need not enter the cell. However, GPCRs activations relate to dynamic events and key intermediate states that arise between the time that a ligand binds and when the G protein is activated. Capturing conformational dynamics of GPCRs and description of intermediate states and its role in activation and signaling has been a formidable challenge, which has hampered progress in understanding activation mechanisms of GPCRs.
Using Fluorine-nuclear magnetic resonance (19F-NMR), mathematical rigidity theory, and molecular dynamics simulations, the international research team has discovered the key mechanism of activation of the human adenosine A2A receptor (A2AR) as it proceeds through signaling pathway. A2AR (also known as caffeine receptor as it is deactivated by caffeine) is a well-known GPCR that is distributed in the nervous system, platelets, immune cells, lungs, heart, and the vasculature. A2AR drugs have been developed to address wound healing, vascular diseases, including atherosclerosis, restenosis, and platelet activation, in addition to inflammation and cancer. Thus, understanding its functional states associated with receptor signaling can yield new opportunities in pharmacology and general understanding of GPCR activation mechanisms.
The researchers focused on biasing key conformational states of A2AR by complexing it with G protein and different ligands to better understand signal transduction and receptor activation. F-NMR showed that A2AR is represented by at least two inactive conformations and three active conformations associated with signaling pathway whose populations are dependent on ligands engagement and G protein interactions. Research team also used Molecular Dynamics simulations to create a structure of A2AR bound to heterotrimer G-protein complex (performed by Kitao lab), where rigidity theory methods of Sljoka identified an activation pathway where A2AR initiates communication with the G-protein which traverses the receptor's ligand binding site and G-protein. Gβγ subunit was discovered to serve as a critical domain for facilitating signaling and activation. Understanding the activation mechanism and functional states of A2AR signaling may provide new opportunities for drug discovery.
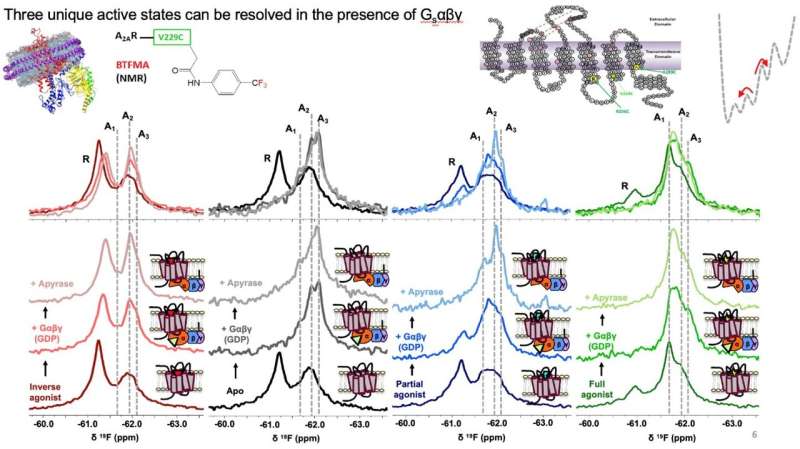
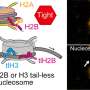
19F-NMR spectra identifies at least 3 active conformational states (A1, A2, A3), and an inactive signature, labeled as R above. Binding of receptor to full heterotrimeric G-protein, Gαβγ, significantly increase the population of active states. The receptor adopts a specific active state A1 when it binds both full agonist and G-protein. Credit: Cell
While the current study provides unprecedented resolution of key functional states associated with receptor signaling, future studies will no doubt focus on other key domains, providing a more comprehensive picture of the activation process.
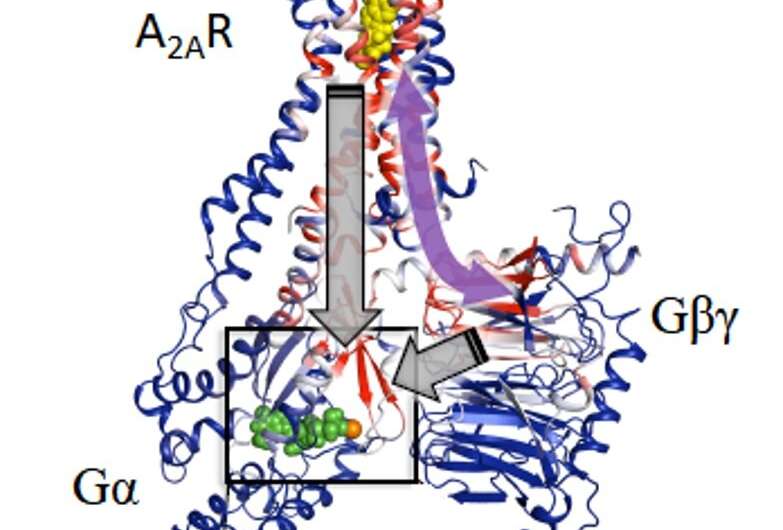
Computational analysis based on modelling and mathematical rigidity theory identifies an activation network pathway encompassing the receptor A2AR, the Gβγ subunit, and the Gα subunits of G-protein, harboring the nucleotide (shown in green). Gβγ subunit was discovered to be critical for signalling and sampling of fully active states of receptor resulting in optimal receptor activation. Credit: Cell
The study is published in the journal Cell.
More information: Cell (2021). DOI: 10.1016/j.cell.2021.02.041
Journal information: Cell
Provided by Tokyo Institute of Technology