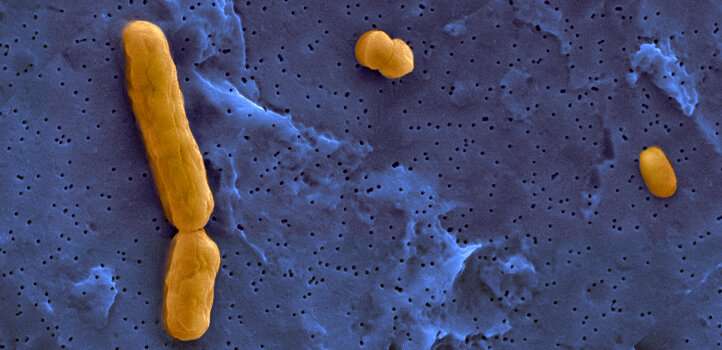
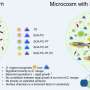
Multiple strains of bacteria exist within an ecosystem: shown here are three bacterial members of the synthetic community viewed under a scanning electron microscope. Credit: 2019 KAUSTReproduced under a Creative Commons license 4.0 from reference 1
A controlled, laboratory approach, along with computer simulations, has helped KAUST researchers to show that bacterial communities can homogenously disperse within aquatic ecosystems, even with slow-flowing water and the persistence of their preferred localized conditions.
"Bacterial communities are a fundamental part of water-based ecosystems, such as oceans, lakes and rivers," explains microbial ecologist Stilianos Fodelianakis. "They represent most of an ecosystem's biomass, and drive carbon and nutrient cycles. If we can understand how these communities change, we might be able to predict changes in the whole ecosystem in light of climate change."
Diverse metacommunities of multiple strains of bacteria exist within an ecosystem, localizing and/or mixing according to the conditions that encourage their growth. These conditions include specific temperature ranges or degrees of acidity or salinity; and the natural movement of the system, such as water flow.
Ecologists have long questioned what exactly causes these metacommunities to homogenize, achieving a state in which bacterial communities across the ecosystem have similar compositions and structures. The assumption was that homogenization occurs when the physical movement within the system is very high, as in fast-flowing rivers, causing high rates of bacterial dispersal and physical mixing of the mediums in which they live. But what favors bacterial metacommunity homogenization more: high rates of bacterial dispersal, or homogenizing environments? And how fast does a system need to move for homogenization to occur?
"We found that homogenization occurs at the high dispersal rates expected in the literature, the so-called 'mass transfer' events, but that it can also happen even at low dispersal rates," says Fodelianakis.
Led by bioscientist Daniele Daffonchio, the team developed a laboratory experiment that enabled them to overcome the challenge of studying changes over time in bacterial communities in their natural environments.
They chose three strains of bacteria that look quite different, making them easy to distinguish from one another, and that grow best at different temperatures. They mixed similar numbers of each strain together, creating multiple, identical communities. One of these communities was placed in each of three linked flasks: the medium could flow between the flasks at four different speeds. The medium in each flask was set at a temperature that favored the growth of one of the three strains.
They found that homogenization of the bacterial communities occurred at all but the slowest speed, even when the temperatures in the flasks changed only slightly. They then conducted 100 different flow rate computer simulations based on their data, focusing particularly on the changes with the two slowest flow rates. This allowed them to identify the precise flow rate (3.85 microlitres per second) at which homogenization occurred within each of the three flasks.
The team is now working on a similar experimental approach to answer other questions in microbial ecology, such as how to quantify the effects of random changes on bacterial populations over time.
More information: Stilianos Fodelianakis et al, Dispersal homogenizes communities via immigration even at low rates in a simplified synthetic bacterial metacommunity, Nature Communications (2019). DOI: 10.1038/s41467-019-09306-7
Journal information: Nature Communications