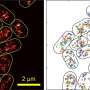
Can bacteria colonies be good neighbours?
A team of German scientists have gained new insights into the way bacteria colonies interact.
The researchers, from the Center for Nanoscience at Ludwig Maximilian University of Munich (LMU), used experiments and computer models to study the population dynamics of bacterial ecological systems.
Their study, which focused on different strains of the E. coli bacteria, publishes today in Physical Biology.
Lead author Dr. Madeleine Opitz, from LMU, said: "Bacterial communities have significant impact on global climate, ecosystems, food chains, and human health. The interactions between constituent bacteria determine the community composition and population dynamics.
"Understanding in detail how this works could help develop new treatments for bacterial infections, for example.
"We have limited understanding of the governing interactions in bacterial communities. Due to the complexity of bacterial interactions, mechanisms controlling ecosystem dynamics are often studied theoretically. Detailed experimental studies supporting these theoretical conclusions are often lacking."
Dr. Opitz and her colleagues studied mixed communities of heterogeneously toxin producing (C strain), toxin sensitive (S strain) and toxin resistant (R strain) bacteria.
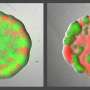
They used multi-scale fluorescence microscopy, which is useful in identifying unknown or underestimated random processes (stochastic effects) in bacterial competition. This allowed them to investigate the microscopic interaction of the R and C strain and to quantify the role of stochastic effects in the spatially structured C-R-S interaction.
The multi-scale time-lapse data enabled the researchers to disentangle the influence of local and global effects of interactions on the macroscopic competition outcome.
They also investigated cheating of the resistant strain, which benefits from the toxin produced by the C strain without paying for the production cost, and determined a distance over which this cheating is maintained.
Dr. Opitz said: "Our results show the early colony patterning is subject to random processes in positioning and toxin production. Then, in a second competition phase, mainly deterministic processes such as bacterial growth and global toxin action determine the following population dynamics."
Together with environmental factors, the variables characterising the early phase predicted the competition outcome reliably.
The team also found that interactions of neighbouring R and C-clusters could amplify local variation, influencing competition dynamics and competition outcome.
Dr. Opitz added: "For this neighbour interaction, we identified cheating of R as a combined action of C and R on S. Analysing the interaction locally on a microscopic scale, we estimated the distance at which the neighbour interaction significantly changes to be around 250 μm."
More information: Benedikt von Bronk et al. Locality of interactions in three-strain bacterial competition in E. coli, Physical Biology (2018). DOI: 10.1088/1478-3975/aae671
Journal information: Physical Biology
Provided by Institute of Physics