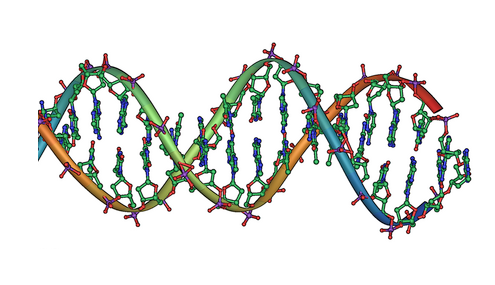
DNA double helix. Credit: public domain
Ever since scientists completed mapping the entire human genome in 2003, the field of DNA sequencing has seen an influx of new methods and technologies designed to help scientists in their search for genetic clues to the evolution of disease and other biological mysteries.
The machines being used are becoming smaller, faster and cheaper. The process of chopping up and reassembling strands of DNA, however, is still far from error proof.
And when studying the one gene most all bacteria have in common, the 16S rRNA gene, getting an accurate portrait is a delicate process.
"It's kind of the elephant in the room when it comes to 16S," said J. Paul Brooks, Ph.D., associate professor in the Department of Statistical Sciences and Operations Research in the Virginia Commonwealth University College of Humanities and Sciences. "Scientists know the accuracy problem exists, but they don't like to talk about it."
Brooks and fellow researchers are talking about it and this spring published in the BMC Microbiology journal one of the most widely read papers in the journal in recent memory.
"The Truth about Metagenomics: Quantifying and Counteracting Bias in 16S rRNA Studies" describes a process that combines experimentation and statistics to account for and reduce bias in DNA analysis.
"The technology develops so fast, and people move on to the next thing without figuring out what they just saw with the last thing," Brooks said. "We're just trying to look at the technologies with clear eyes."
Brooks is a member of the VCU Vaginal Microbiome Consortium, a group of researchers focused on women's health. As such, the recent study on bias looked at seven strains of bacteria critical to the vaginal microbiome, another word for the community of microscopic organisms present in an environment. These organisms mostly perform necessary biological functions, but some can be pathogenic.
This is why scientists take DNA from samples and sequence it to see if they can identify mutations or other genetic activity that could lead to disease. Studying bacteria in the vaginal microbiome will help researchers better understand premature birth, sexually transmitted diseases and other women's health issues.
Brooks and company looked at the entire process involved: extracting DNA from a sample, amplifying it and then sequencing and classifying it. These steps all affect bacteria in different ways and contribute to bias, Brooks said.
For instance, some bacteria yield DNA easier than others during extraction.
With amplification, scientists use a process called polymerase chain reaction, a quick and automated way to make copies of many DNA segments. There is a temptation to do fewer cycles to reduce the chances of bias, but that could mean missing a rarer strain of bacteria.
For the recent study, researchers used mixtures of samples containing different amounts of bacteria. The goal was to put these mixtures through the usual processing pipeline to see if what comes out matches what went in. It is a balance of concrete truth and observation.
Researchers start with an input, in this case the proportions of bacteria in a given mixture. This is the truth. Using the observed data, statistical models are constructed to predict the proportion of bacteria in a sample after going through the sequencing process.
"We can also use the data to create inverse models," said David J. Edwards, Ph.D., also an associate professor in the Department of Statistical Sciences and Operations and co-author of the paper. "That is, we try to go the other way. In other words, can we take what bacterial proportions were actually observed in order to model the truth?"
The models are based on mixture experiments commonly used in the chemical and process industries, the kind used to determine formulations for gasoline, paint or even wine.
"Let's say you're baking cookies. You'd have to mix together differing amounts of ingredients like flour, milk, butter, eggs and chocolate chips," said Edwards said. "To come up with the optimal recipe for cookie dough, you might conduct an experiment with these ingredients by mixing them in different proportions. Obviously, cookie dough made with 100 percent milk isn't going to do anything for us. And I doubt that one-third flour, one-third milk and one-third butter would taste very good either."
The models give researchers a better idea of bacteria present in an entire community based on a small sample, which by itself could distort the bigger picture.
"What you observe in your sample is not always an accurate picture of what's actually there," Brooks said.
There were cases where a sample was created with 50 percent of a certain bacteria and 50 percent of another. But when run through the new process, the split was closer to 85 percent and 15 percent.
The pitfalls still occur even when scientists make use of a variety of extraction kits to compensate for bias.
"If you have a pool of DNA and you go fishing for that 16S gene, the bait you use is going to influence the process," Brooks said. "No matter what method you choose, there is going to be bias."
The title of the recently published paper has caused a small firestorm on Twitter.
The truth about metagenomics is that metagenomics refers to the study of genetic material taken from the environment. The genome is the entire genetic makeup of an organism, and Brooks' team focused their study on the lone 16S rRNA gene.
"Is it a provocative title? Of course," Brooks said. "But by targeting one gene, we can figure out what genomes are there."
Researchers are pushing ahead with their models, which were designed for the seven bacteria analyzed in the recent study. They hope to conduct additional experiments to develop models that can be applied to any environment and any bacteria.
"We have questions we want to answer," Brooks said. "My dream would be to have this universal quality control that can be used for reproducible research. If we're going to do that, though, we need to make sure the community compositions that we are observing reflect what is actually in the environment."
More information: "The truth about metagenomics: quantifying and counteracting bias in 16S rRNA studies." BMC Microbiology 2015, 15:66 DOI: 10.1186/s12866-015-0351-6
Journal information: BMC Microbiology
Provided by Virginia Commonwealth University