Naomi Marty
(PhysOrg.com) -- University of Arkansas researchers have characterized a membrane receptor protein and its binding mechanism from chloroplasts in plants and determined that it shares a commonly shaped binding site and mechanism with a similar protein found in E. coli.
The paper, published in the Journal of Biological Chemistry, was chosen as the research paper of the week for its significance and overall importance to the field.
“It’s strange to think of the processes in plants having similarities to E. coli, but they do,” said Robyn Goforth, research professor of biological sciences.
Researchers are still learning how proteins get from where they are manufactured to where they do their work. Goforth and graduate student Naomi Marty examined the path of a particular protein in plants that shepherds light-harvesting chloroplast proteins into the thylakoid membrane. Although bacteria do not have chloroplasts, they do have a similar mechanism by which proteins get transported from one location to another through the cytosolic membrane.
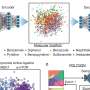
Goforth and Marty looked at the signal recognition particle pathway in plants, which is responsible for taking light-harvesting proteins from where they are made to where they are used. They identified the binding mechanism for the signal recognition particle receptor, a membrane-binding protein that helps bring the light harvesting chloroplast protein to the membrane and allows it to bind there.
To do this, they isolated chloroplast membranes from pea plants, then introduced the modified receptor, first taking off three amino acids, then six, then nine. They then examined the modified proteins’ ability to move light-harvesting proteins to the membrane. As a result, they were able to identify an 18-amino acid region that is essential to the protein transport process and that changes structure when interacting with the membrane. They identified two phenylalanine residues, found in the receptor proteins in both plants and bacteria, that prove essential to the signal recognition particle receptor’s role in binding proteins to the membrane.
Together with colleagues in the department of biological sciences and the department of chemistry and biochemistry, they examined the structure of the protein when it interacts in the membrane and in solution. They found that this region of the receptor protein had different structures in the two different environments.
“When you change the phenylalanine, you don't get the structural switch,” Goforth said. “This peptide is both necessary and sufficient for targeting proteins to the membrane.”
They also studied a similar transportation pathway found in E. coli, whereby certain proteins are taken to the membrane to act as exterior sensors.
“What we show here is that both the E. coli and the chloroplast receptor proteins react the same way at the membrane,” Marty said.
The team consisted of Goforth, Marty, Alicia Kight, Nathaniel Lewis, Daniel Fologea and professor Ralph Henry of the department of biological sciences and Dakshinamurthy Rajalingam and professor Suresh Kumar of the department of chemistry and biochemistry. All are researchers in the Center for Protein Structure and Function in the J. William Fulbright College of Arts and Sciences.
Provided by University of Arkansas (news : web)