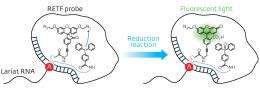
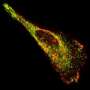
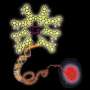
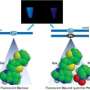
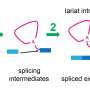
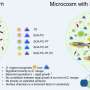
Figure 1: Schematic representation of the reduction reaction between the dye precursor and triphenyl phosphine within the lasso structure of an LaRNA molecule. When these probes meet at the junction (A), fluorescent light is emitted, effectively tagging the molecule. Credit: Reproduced in part, with permission, from Ref.1 © 2012 Wiley-VCH
Numerous biological processes depend on molecules called lariat RNAs (LaRNAs). These lasso-shaped structures form in the cell during RNA splicing. During this process, transcribed RNA strands convert to messenger RNA before undergoing translation into proteins. A way to quickly and efficiently characterize these molecules in living cells is now available, thanks to a method developed by a research team led by Hiroshi Abe of the RIKEN Advanced Science Institute, Wako. The method identifies LaRNAs using molecular pairs called reduction-triggered fluorescent (REFT) probes. When in close proximity to LaRNAs, these probes react and generate a fluorescent signal.
In contrast to earlier approaches to identify elusive LaRNAs, Abe and his team adopted a simple, non-enzymatic strategy. They exploited RNA strands to make a template of a reaction between the probe components, consisting of a fluorescent dye precursor and a reducing agent called triphenyl phosphine (Fig. 1). To position the probes, they attached the precursor and the triphenyl phosphine to separate DNA strands and then paired them with the complementary RNA template. In an open linear template, the REFT components were too far apart to react. However, the circular loop of LaRNA templates brought the reacting species close enough to facilitate the fluorescence-inducing reaction.
Performance assessments using these probe-linked DNA strands, which could bind different portions of the LaRNAs, showed that a fluorescent signal was emitted only when the reacting components could access the target at the lasso junction. “This offers a high signal-to-background ratio during the detection,” says Abe. His team also demonstrated that bigger LaRNA loops increased the accessibility of the probes, enhancing the fluorescent response. Furthermore, splicing reactions carried out in vitro revealed that the probes did not hinder the function of associated catalysts and LaRNA production in cells.
Abe and colleagues compared their REFT system with the conventional fluorescence resonance energy transfer (FRET) approach, which also relies on probes that fluoresce when next to each other. Without any optimization, the REFT probes outperformed the FRET probes in distinguishing LaRNAs from similar structures. “Our probes recognize the unusual secondary structure of the [lassos] whereas conventional methods can only be applied to linear RNA structures,” explains Abe. He adds that the flexible RNA molecules can adopt various geometries that strongly resemble the lariat architecture causing FRET to produce false-positive results.
The team is currently planning to use the REFT probes to image RNA. “An important application of our system is the real-time monitoring of splicing in living cells,” says Abe.
More information: Furukawa, K., et al. Fluorescence detection of intron lariat RNA with reduction-triggered fluorescent probes. Angewandte Chemie International Edition 50, 12020–12023 (2011).
Journal information: Angewandte Chemie International Edition
Provided by RIKEN