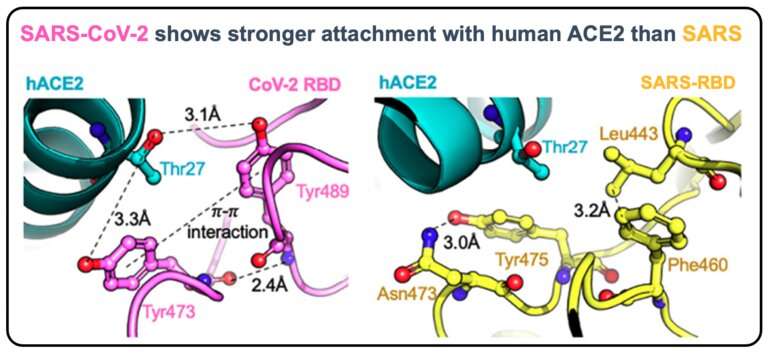
This schematic shows the difference in interactions between SARS-CoV and human ACE2 versus SARS-CoV-2 and human ACE2. Credit: Ratul Chowdhury, Penn State
The binding of a SARS-CoV-2 virus surface protein spike—a projection from the spherical virus particle—to the human cell surface protein ACE2 is the first step to infection that may lead to COVID-19 disease. Penn State researchers computationally assessed how changes to the virus spike makeup can affect binding with ACE2 and compared results to those of the original SARS-CoV virus (SARS).
The researchers' original manuscript preprint, made available online in March, was among the first to computationally investigate SARS-CoV-2's high affinity, or tendency to bind, with human ACE2. The paper was published online on Sept. 18 in the Computational and Structural Biotechnology Journal. The work was conceived and led by Costas Maranas, Donald B. Broughton Professor in the Department of Chemical Engineering, and his former graduate student Ratul Chowdhury, who is currently a postdoctoral fellow at Harvard Medical School.
"We were interested in answering two important questions," said Veda Sheersh Boorla, doctoral student in chemical engineering and co-author on the paper. "We wanted to first discern key structural changes that give COVID-19 a higher affinity towards human ACE2 proteins when compared with SARS, and then assess its potential affinity to livestock or other animal ACE2 proteins."
The researchers computationally modeled the attachment of SARS-CoV-2 protein spike to ACE2, which is located in the upper respiratory tract and serves as the entry point for other coronaviruses, including SARS. The team used a molecular modeling approach to compute the binding strength and interactions of the viral protein's attachment to ACE2.
The team found that the SARS-CoV-2 spike protein is highly optimized to bind with human ACE2. Simulations of viral attachment to homologous ACE2 proteins of bats, cattle, chickens, horses, felines and canines showed the highest affinity for bats and human ACE2, with lower values of affinity for cats, horses, dogs, cattle and chickens, according to Chowdhury.
"Beyond explaining the molecular mechanism of binding with ACE2, we also explored changes in the virus spike that could change its affinity with human ACE2," said Chowdhury, who earned his doctorate in chemical engineering at Penn State in fall 2019.
Understanding the binding behavior of the virus spike with ACE2 and the virus tolerance of these structural spike changes could inform future research on vaccine durability and the potential for the virus to spread to other species.
"The computational workflow that we have established should be able to handle other receptor binding-mediated entry mechanisms for other viruses that may arise in the future," Chowdhury said.
More information: Ratul Chowdhury et al, Computational biophysical characterization of the SARS-CoV-2 spike protein binding with the ACE2 receptor and implications for infectivity, Computational and Structural Biotechnology Journal (2020). DOI: 10.1016/j.csbj.2020.09.019
Provided by Pennsylvania State University