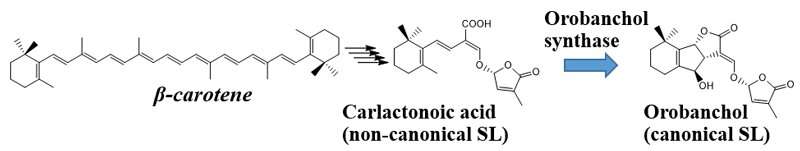
Figure 1: Diagram showing the biosynthesis of orobanchol from β-carotene. The orobanchol synthase indicated by the blue arrow, which provided the reaction catalyst, was illuminated by this research. Credit: Kobe University
Strigolactones (SLs) are a class of chemical compounds found in plants that have roles as plant hormones and rhizosphere signaling molecules. They regulate plant architecture and promote germination of root parasitic weeds that have detrimental effects on plant growth and production.
A study was conducted as part of the SATREPS (Science and Technology Research Partnership for Sustainable Development) program by Dr. Wakabayashi, Prof. Sugimoto and their colleagues at the Graduate School of Agricultural Science, Kobe University, in collaboration with researchers from the University of Tokyo and Tokushima University. They discovered the orobanchol synthase responsible for converting the SL carlactonoic acid, which promotes symbiotic relationships with fungi, into the SL orobanchol, which causes root parasitic weeds to germinate.
By knocking out the orobanchol synthase gene using genome editing, they succeeded in artificially regulating SL production. This discovery will lead to greater understanding of the functions of each SL and enable the practical application of SLs in the improvement of plant production.
The results of this study were published in Science Advances on December 18, 2019.
Strigolactones (SL) are a class of chemical compounds that were initially characterized as germination stimulants for root parasitic weeds. SLs have also received attention for their other functions. They play an important role in controlling tiller bud outgrowth and also in promoting mycorrhizal symbiosis in many land plants, whereby plants and fungi mutually exchange nutrients.
Figure 2: Tomato plant branching. (Red arrows indicate auxiliary buds). Wild type (left), SlCYP722C-KO (center), SL deficient mutant (Slccd8, right). Credit: Kobe University
Up until now, around 20 SLs have been isolated; with differences in stereochemistry in the C ring and modifications in the A and/or B rings. In recent years, SLs with unclosed BC rings have been discovered. Currently, SLs with a closed ABC ring are designated as canonical SLs, whereas SLs with an unclosed BC ring are non-canonical SLs. However, it is not clear which compounds function as hormones and which compounds function as rhizosphere signals.
If SL production could be suppressed, plants would induce the germination of fewer root parasitic weeds and their adverse effects on crop production would be mitigated. By increasing SL production, on the other hand, plant nutrition would be improved through the promotion of relationships with mycorrhizal fungi. Furthermore, manipulation of the endogenous levels of SL would control plant architecture above ground. Understanding the functions of individual SLs would lead to the development of technology to artificially control plant architecture and the rhizosphere environment. Consequently, there is much interest in how these SLs are biosynthesized.
It has been elucidated that SLs are biosynthesized from β-carotene. Four enzymes are involved in conversion of β-carotene to carlactonoic acid (CLA), a common intermediate of SL biosynthesis. In Japonica rice, conversion of CLA into orobanchol proceeds with two enzymes catalyzing two distinct steps. However, the biosynthesis pathway for orobanchol in other plants remained unknown. This study discovered the novel orobanchol synthase, which converts CLA into orobanchol in cowpea and tomato plants (Figure 1).
The researchers had isolated orobanchol from cowpea root exudates and determined the structure. From metabolic experiments using cowpea, it was predicted that cytochrome P450 would be involved in the conversion of CLA into orobanchol. In this study, cowpea plants were grown in phosphate rich and poor conditions, where orobanchol production was restricted and promoted, respectively. The genes expressed in the roots of plants in both conditions were comprehensively compared. The group screened for CYP genes whose expression correlated with orobanchol production, expressed them as recombinant proteins, and performed an enzyme reaction assay.
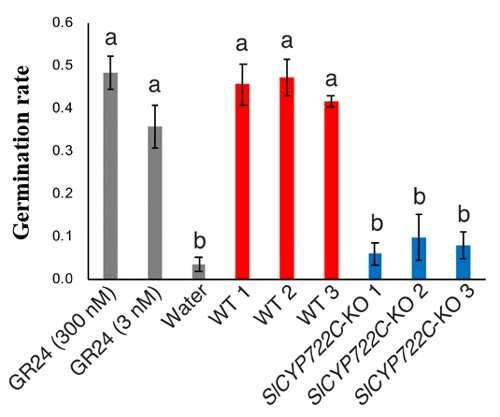
Figure 3: Germination rate of the root parasitic weed p.aegyptiaca in the hydroponic medium of SlCYP722C-KO mutants Levels are far lower compared to the wild type. Credit: Kobe University
From these results, it was understood that the VuCYP722C enzyme catalyzed the conversion of CLA to orobanchol. Furthermore, the SlCYP722C gene, a homolog of VuCYP722C in tomato was confirmed to be an orobanchol synthase gene. The SlCYP722C gene was knocked out (KO) in tomato plants using genome editing. In contrast to the wild type (control) tomato plants, orobanchol was not detected in root exudates of the KO plants, with CLA being detected instead.
Thus, the research group proved that SlCYP722C is the orobanchol synthase in tomato that converts the non-canonical SL CLA into the canonical SL orobanchol. The architecture of the KO and wild-type plants was comparable (Figure 2). This demonstrated that orobanchol doesn't control plant architecture in tomato plants. It is thought that these KO tomato plants would still be able to benefit from mycorrhizal fungi, as the activity of CLA against the hyphal branching of the fungi was comparable with that of canonical SLs. Furthermore, it was found that the germination rate of the root parasitic weed Phelipanche aegyptiaca was significantly lower in the hydroponic media of the KO tomato plants (Figure 2). P. aegyptiaca causes great damage to tomato production all over the world, especially around the Mediterranean region. This research showed that it is possible to limit the damage that this parasitic weed does to tomato production by knocking out the orobanchol synthase gene.
The researchers succeeded in preventing the synthesis of the major canonical SL orobanchol and accumulating the non-canonical SL carlactonoic acid. The same method can be utilized to elucidate the genes responsible for the biosynthesis of other canonical SLs. Further understanding of the functions of various SLs would allow plants to be manipulated in order to maximize their performance under adverse cultural conditions. Root parasitic weeds detrimentally affect not only tomato but a wide range of other crops including species of Solanaceae, Leguminoceae, Cucurbitaceae and Poaceae. These results will lead to the development of research to alleviate the damage inflicted by root parasitic weeds and increase food production worldwide.
More information: Takatoshi Wakabayashi et al. Direct conversion of carlactonoic acid to orobanchol by cytochrome P450 CYP722C in strigolactone biosynthesis, Science Advances (2019). DOI: 10.1126/sciadv.aax9067
Journal information: Science Advances
Provided by Kobe University