Douglas Rees (left) and Garnet Chan (right). Credit: Caltech
Nitrogenases, the enzymes responsible for converting atmospheric nitrogen into the nitrogen-based compounds used by living things, have been a major focus of chemistry research for decades because of the pivotal role that nitrogen fertilizers play in feeding the planet's burgeoning population.
Two Caltech researchers have been at the forefront of studies of the molecular structure of these important enzymes: Douglas Rees and Garnet Chan.
Rees, Caltech's Roscoe Gilkey Dickinson Professor of Chemistry, investigator with the Howard Hughes Medical Institute, and dean of graduate studies, is one of the pioneers of this research. In 1992, Rees published the first paper describing the structure of a nitrogenase enzyme. Since then, his lab has continued to investigate the structure and behavior of these molecules. In recognition of his work, the Royal Swedish Academy of Sciences in September named Rees as one of the winners of this year's Gregori Aminoff Prize. The prize is awarded annually to researchers for their accomplishments in the field of crystallography.
"The mechanism of biological nitrogen fixation has intrigued chemists for over a century," Rees says. "My group has approached this question by determining the X-ray crystal structures of the nitrogenase proteins that catalyze the conversion of atmospheric dinitrogen to ammonia under physiological conditions. Through the efforts of a remarkable group of graduate students and postdoctoral fellows, in a continuing collaboration with my former postdoctoral advisor, James Howard, we have been able to define the molecular architecture of nitrogenase in exquisite detail, with particular focus on the unusual metalloclusters that provide the active site for this reaction."
Chan, Caltech's Bren Professor of Chemistry, has also been researching the structure of nitrogenases, using computational methods that offer complementary information to the studies by Rees. Chan's specialty is quantum chemistry, a field that attempts to explain chemical properties through principles of quantum mechanics.
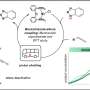
This month, Chan and his coworkers published a new paper on nitrogenase in Nature Chemistry. In that paper, they outline the electronic structure—the arrangement of electrons—of the so-called P-clusters found in nitrogenase enzymes, as determined through computer simulations. P-clusters are regions within the enzyme that consist of several iron and sulfur atoms bonded together. They are responsible for part of the chemical process that turns nitrogen gas into nitrogen compounds like ammonia, or nitrogen fixation.
Determining how electrons are arranged in these P-clusters would help researchers better understand the mechanism that underlies nitrogen fixation, but these arrangements have proven difficult to study. Chan's work relies on new computational techniques developed in his lab based on the ideas of quantum information theory.
"Without Doug's seminal studies we would not know how the atoms are arranged in the P-cluster," Chan says. "But with the knowledge of these positions, we can now use theoretical quantum mechanics to visualize where the electrons are, adding a new chapter to this ongoing story."
Chan's paper, titled, "Electronic landscape of the P-cluster of nitrogenase as revealed through quantum many-electron wavefunction simulations," appears in the September 30 issue of Nature Chemistry.
More information: Zhendong Li et al. Electronic landscape of the P-cluster of nitrogenase as revealed through many-electron quantum wavefunction simulations, Nature Chemistry (2019). DOI: 10.1038/s41557-019-0337-3
Journal information: Nature Chemistry
Provided by California Institute of Technology