Nucleosides are fundamental building blocks of genetic material which makes them attractive for a number of biologically relevant applications and as potential pharmaceuticals. At The City College of New York, scientists are developing facile methods for modifying nucleoside structures to make chemical processes more efficient.
Mahesh Lakshman, professor in City College's Department of Chemistry and Biochemistry, leads a team that has developed easy access to relatively complex nucleoside analogues.
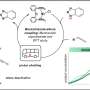
They have devised a carbon-nitrogen bond forming strategy leading to new nucleoside analogues, using "hypervalent iodine" reagents that are of increasing importance in molecule building. However, the nucleosides used had several possible centers that could combine in these reactions.
The researchers found that these nucleosides bend into certain patterns to drive specific reactive centers to combine, depending upon the conditions used. They also analyzed the possible fundamental pathways of these reactions.
Their work was ranked "highly important" in the review process and was the cover feature in issue 21 of the journal ChemCatChem.
The research expands on previous modifications of nucleosides conducted by Lakshman, a Fellow of the Royal Society of Chemistry. Working with postdoctoral associate Sakilam Satishkumar, he has sought to manipulate nucleosides using hypervalent iodine reagents, in order to access new molecules, which showed interesting fluorescence properties depending upon structure.
Because they are genetic building materials found in all living organisms, nucleosides possess great potential, from drug candidates to detection tools.
More information: Sakilam Satishkumar et al. Pd-Catalyzed versus Uncatalyzed, PhI(OAc)2 -Mediated Cyclization Reactions of N 6 -([1,1′-Biaryl]-2-yl)Adenine Nucleosides, ChemCatChem (2017). DOI: 10.1002/cctc.201700918
Provided by City College of New York