Credit: New York University
At the first hints of a disease outbreak, epidemiologists, health care providers, policy makers, and scientists turn to sophisticated predictive models to determine how an illness is spreading and what should be done to minimize contagion. A research collaboration between the New York University Tandon School of Engineering and Politecnico di Torino in Italy is upending the traditional modeling process, yielding predictions that are both simpler to calculate and more attuned to a hyper-connected world.
All predictive models correlate the movement of an illness through a population over time, but current simulations fail to account for a seemingly obvious idea: that mobility and activity varies among people, and that these variations impact the likelihood of contracting or spreading an illness.
A new paradigm was explained in a paper published in Physical Review Letters by Maurizio Porfiri, a professor of mechanical and aerospace engineering at NYU Tandon, Alessandro Rizzo, a visiting professor at NYU Tandon and an associate professor of control engineering at Politecnico, and Lorenzo Zino, a Politecnico doctoral student in pure and applied mathematics.
The researchers assume that some people are more active, some less so, and their model accounts for how these differences may impact disease spread. Their approach permits nuanced modeling of different illnesses—from a highly contagious airborne virus such as influenza, which moves quickly among people with high mobility but is limited by those who seclude themselves, to a virus like HIV, which has a long latency period and slower transmission rate.
"The way I move is the way I catch a disease," said Porfiri. "We're changing the point of view from which we start outbreak simulations because we can't understand how a small outbreak evolves into an epidemic without understanding how different people's activity levels help spread it."
Several traditional models assume homogeneity within the community. "It's like sick people are all in a specific place, connecting with a set number of people, and that's not realistic," said Rizzo. "Some people make more connections than others, and the scale of those connections may be comparable to the scale of the disease."
Porfiri and Rizzo explained that traditional simulations use a "discrete time/continuous activity" approach, which typically requires extensive and lengthy simulations. The researchers employ simpler systems of coupled differential equations that allow for the manipulation of factors that can influence disease spread.
This is the first piece of research to emerge from a three-year, $375,000 National Science Foundation grant awarded to the team to study the concurrent evolution of the dynamics of infectious diseases and the networks through which they spread. The research was also funded in part by grants from the U.S. Army Research Office (ARO) and Compagnia di San Paolo.
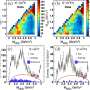
The team has developed one of the few disease modeling approaches that uses heterogeneities in activity levels as a factor in spreading disease. In experiments to test their model, the team successfully predicted the movement of influenza on a university campus and the spread of a trending topic on Twitter.
"We have infinite possibilities to see the impact of interventions," said Porfiri. "We can understand how vaccines, quarantine, or other parameters influence contagion. Some illnesses catch fire, while others are quashed immediately. This framework allows for analysis of why and how that happens."
In the future, the researchers expect that this model will aid management efforts during an outbreak, including implementing vaccination strategies, evaluating the risks and benefits of travel bans, and gauging the effectiveness of disease prevention campaigns.
More information: Lorenzo Zino et al. Continuous-Time Discrete-Distribution Theory for Activity-Driven Networks, Physical Review Letters (2016). DOI: 10.1103/PhysRevLett.117.228302
Journal information: Physical Review Letters
Provided by New York University