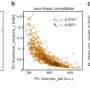
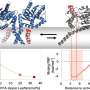
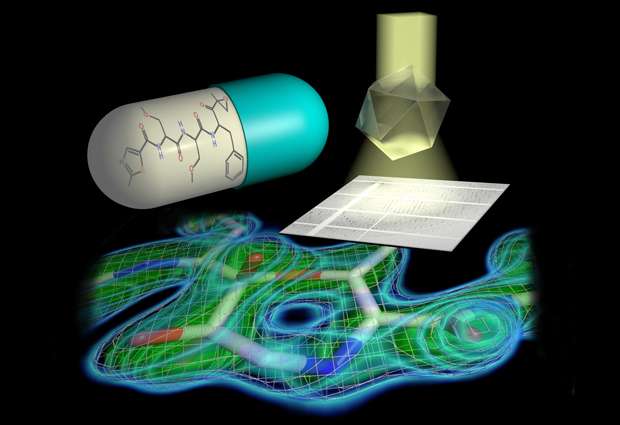
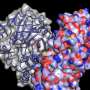
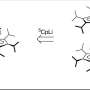
Tailored parallel X-rays perfectly matching the dimensions of the protein crystals enabled the scientists to determine the proteasome structure in unprecedented detail (top right). They could show that a 7-ring is formed by the reaction of the inhibitor with the proteasome (bottom). This knowledge paves the way for the design of new therapeutics. Credit: Hartmut Sebesse / Max Planck Institute for Biophysical Chemistry
Cancer cells are more dependent on a cellular garbage disposal unit—the proteasome—than healthy cells, and cancer therapies take advantage of this dependency. Scientists at EMBL and MPI determined the proteasome's 3D structure in unprecedented detail and have deciphered the exact mechanism by which inhibitor drugs block the proteasome. Their surprising results, published in Science, will pave the way to develop more effective treatments.
Malignant cancer cells not only proliferate faster than most healthy cells in our bodies. They also generate more "junk", such as faulty and damaged proteins. This makes cancer cells inherently more dependent on the most important cellular garbage disposal unit, the proteasome, which degrades defective proteins and removes them from circulation. Treatments for some types of cancer, such as multiple myeloma—a type of bone marrow cancer—exploit this dependence. Patients are treated with inhibitors, which selectively block the proteasome. The ensuing pile-up of cellular junk overwhelms the cancer cell, ultimately killing it. A team of researchers from Göttingen and Hamburg have now succeeded in determining the 3D structure of the human proteasome in unprecedented detail and have deciphered the exact mechanism by which inhibitors block the proteasome.
In order to understand how cellular machines such as the proteasome work, it is essential to determine their three-dimensional structure in detail. With its more than 50.000 atoms, the barrel-shaped proteasome, however, is a true challenge for structural biologists. A group of scientists led by Ashwin Chari at the Max Planck Institute (MPI) for Biophysical Chemistry in Göttingen and Gleb Bourenkov at EMBL have now managed to determine the three-dimensional structure of the human proteasome at an unprecedented resolution of 1.8 Ångström—enabling them to pinpoint the position of single atoms in the garbage disposal unit.
In a next step, the researchers solved the structure of the proteasome bound to four different inhibitors that are either already used in the clinic or are currently undergoing clinical trials. "The substantial improvement in resolution compared to previous proteasome structures has allowed us to establish the exact chemical mechanism by which inhibitors block the proteasome. This knowledge makes it possible to optimize inhibitor design and efficacy—since only inhibitors tailored to the proteasome shut it down completely," says Chari, project group leader in the Department of Structural Dynamics headed by Holger Stark at the MPI for Biophysical Chemistry.
The scientists discovered an important detail in the proteasome's active site. The active site is what enables the proteasome to degrade the cell's junk, and it is what the inhibitor drugs bind to in order to shut off that activity. In contrast to the common perception, a 7-ring structure is formed by the chemical reaction of inhibitor and proteasome active site, which contains an additional so-called methylene group. This has far-reaching consequences for the inhibitor's efficacy and chemical mechanism, the researchers explain. "Even though a methylene group just comprises one carbon atom and its two associated protons amidst the more than 50.000 atoms of the proteasome, it decisively influences which chemical features make the inhibitor most effective in blocking the proteasome," says Thomas Schneider, who leads a group at EMBL. "This has to be taken into account when developing new inhibitors and searching for new drug candidates," adds Holger Stark. The researchers have already filed a patent application for the chemical procedure to design such inhibitors. "Clinical applications are always preceded by knowledge about targets - therefore, the details, where every atom counts, make all the difference," Bourenkov states.
Huge effort reveals a small difference
The project's success is the result of fantastic teamwork, as Max Planck researcher Chari emphasizes: "A group of scientists, all experts in their respective fields, contributed their specialized knowledge, expertise, and complemented each other perfectly." Structural biologists, physicists, enzymologists, and biochemists of the MPI for Biophysical Chemistry, EMBL, and the University of Göttingen developed several innovative procedures.
To determine a molecule's structure using X-ray crystallography, scientists grow crystals of that molecule, then shine a powerful beam of X-ray light on the crystal. Based on how the X-rays scatter after hitting the crystal, researchers can deduce the molecule's 3D structure. Fabian Henneberg and Jil Schrader, junior scientists in Stark's department and first authors of the report now published in Science, used a new method to purify proteasomes and grow the high-quality crystals that made it possible to solve its 3D structure in such detail. The scientists have filed for a second patent application based on the purification and crystallization procedure employed in this work. "The pipeline we use to purify and crystallize the proteasome with and without inhibitors is also suitable to discover new proteasome inhibitors - in an industrial setting, screening several hundred compounds per week could be feasible," Chari predicts.
However, the crystals were only one element of the project's success. The second were the cutting-edge instruments developed by the EMBL research facility on the Deutsches Elektronen Synchrotron (DESY) campus in Hamburg. "The DESY light source generates X-rays of exceptional quality. With the help of powerful X-ray optics, we were able to tailor X-rays to perfectly suit the crystallized proteasome. Only this made it possible to determine the proteasome structure in unprecedented detail," concludes Bourenkov.
More information: "The inhibition mechanism of human 20S proteasomes enables next-generation inhibitor design" Science, science.sciencemag.org/cgi/doi … 1126/science.aaf8993
Journal information: Science
Provided by European Molecular Biology Laboratory