The worst of the recent Ebola epidemic is over, but the threat of future outbreaks lingers. Monitoring the virus requires laboratories with trained personnel, which limits how rapidly tests can be done. Now scientists report in ACS' journal Analytical Chemistry a handheld instrument that detects Ebola quickly and could be used in remote locations.
The gold standard method for identifying the Ebola virus in a blood sample requires packaging samples in cooled containers and sending them to specialized laboratories, often far away from where patients live. Lab personnel use a method called reverse-transcription polymerase chain reaction, or RT-PCR, to check for the virus. The prolonged testing process delays detection, treatment and real-time monitoring of viral loads in body fluids that can harbor the virus even after it's no longer detected in blood. To make reliable and fast Ebola detection more accessible, Pavel Neuzil and colleagues developed a cellphone-sized device, which they reported in a previous paper.
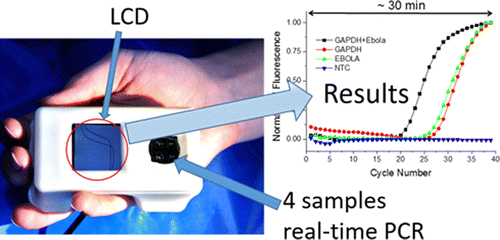
In their latest study, the researchers designed and tested the instrument, which could simultaneously perform four RT-PCRs that included two controls and two patient blood samples. Conventional tests require several hours to more than a day for results to come in. The new process took less than 37 minutes, and the amount of blood required was minute and could potentially come from just a finger prick.
The device successfully detected the Ebola RNA. In addition to diagnosing the illness, the test also yielded information about how many RNA copies each sample contained. In addition to diagnosing the illness, they say that the tool could also potentially help health care workers track patients' viral loads in semen, breast milk and eye fluids after recovery.
More information: Christian D. Ahrberg et al. Palm-Sized Device for Point-of-Care Ebola Detection, Analytical Chemistry (2016). DOI: 10.1021/acs.analchem.6b00278
Abstract
We show the utilization of a recently developed cellphone-sized real-time polymerase chain reaction (PCR) device to detect Ebola virus RNA using single-step reverse transcription PCR (RT-PCR). The device was shown to concurrently perform four PCRs, each with a sample volume of 100 nL: one positive control with both Ebola and GAPDH RNA and one negative control. The last two positions were used to measure the GAPDH and the Ebola content of a sample. A comparison of threshold cycles (CT) from the two samples provided relative quantification. The entire process, which consisted of reverse transcription, PCR amplification, and melting curve analysis (MCA), was conducted in less than 37 min. The next step will be integration with a sample preparation unit to form an integrated sample-to-answer system for point-of-care infectious disease diagnostics.
Journal information: Analytical Chemistry
Provided by American Chemical Society