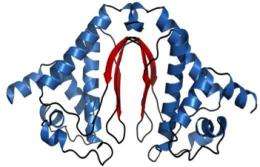
Crystal structure of DUF269 (3NJ2). The β-sheets at the dimer interface, highlighted in red, form the framework for a solvent-accessible cleft.
(Phys.org) -- The crystal structure of a protein in a bacterium being studied for its renewable energy potential suggests that the protein's biological function may be unique to a process leading to hydrogen gas production, according to researchers from Pacific Northwest National Laboratory and Brookhaven National Laboratory. They determined the structure for a protein that falls into a family of proteins with a "Domain of Unknown Function," annotated DUF269, from Cyanothece 51142, a photosynthetic bacterium with a completely different lifestyle in the day and the night.
They found that a solvent-accessible cleft with conserved charged residues at the interface of the asymmetric unit of the crystal (see figure) could be an active site or ligand-binding surface for the protein's biological function. The protein fold observed for the Cyanothece appears to be unique to the DUF269 family of proteins.
By day Cyanothece 51142 uses sunlight to convert carbon dioxide into molecular oxygen and glucose, a process called photosynthesis. At night it uses the glucose to convert atmospheric N2 into biologically available ammonia, a process called nitrogen fixation.
Understanding how this single-celled organism regulates photosynthesis and nitrogen fixation may have big dividends for multi-celled organisms such as man because a byproduct of the nitrogen fixation process is hydrogen gas (H2), an appealing source of renewable and green energy. Clues to understanding the process of regulating nitrogen fixation may rest in the 34 genes present in the nitrogen fixation gene collection of Cyanothece. DUF269 is one of these genes, and to date, belongs to a subset of 16 of the 34 gene products in the nitrogen fixation operon whose function is unknown.
Crystals of cce_0566 were grown at PNNL, harvested, and sent to BNL for X-ray diffraction data collection at the National Synchrotron Light Source. NMR and other biophysical data collected at EMSL confirmed that the biological unit of DUF269 in solution was the same as observed in the asymmetric unit of the crystal, a dimer.
Further biochemical experiments are in progress to determine the biological function of this protein with a fold unique to the DUF269 family of proteins.
More information: GW Buchko and H Robinson. 2012. "Crystal Structure of cce_0566 from Cyanothece 51142, a Protein Associated with Nitrogen Fixation in the DUF269 Family." FEBS Letters 586: 350-355. DOI: 10.1016/j.febslet.2012.01.037
Journal information: FEBS Letters
Provided by Pacific Northwest National Laboratory