Designer enzyme uses unnatural amino acid for catalysis
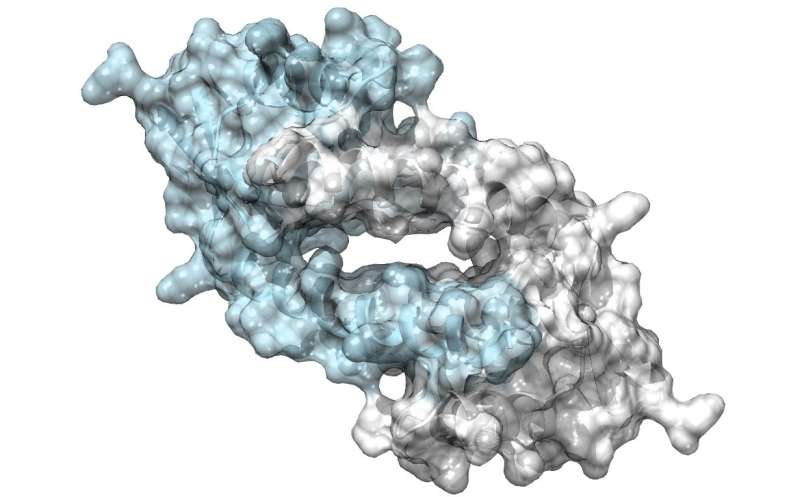
University of Groningen chemists have created a new enzyme with an unnatural amino acid as its active centre. They made the enzyme by modifying an antibiotic binding protein which normally acts as a bacterial transcription factor. Further modifications of the reactive site can create different enzymes for use in chemical synthesis. A description of the new enzyme was published in Nature Chemistry on 2 July.
The chemical industry is working hard to replace conventional processes, which can use toxic chemicals or a lot of energy, with more environmentally friendly alternatives. A popular alternative is enzymes, which usually function in water at moderate temperatures. However, the number of reactions catalysed by natural enzymes is limited compared with what is available in chemistry. "That is why enzyme design has really taken off in recent years," says University of Groningen Professor of Biomolecular Chemistry Gerard Roelfes.
Increased catalysis
Enzymes can be engineered in different ways. Yet, so far, only the 20 naturally occurring amino acids have been used to create the active site of enzymes. This limits the number of reactions these designed enzymes can perform, argues Roelfes: "We think that incorporating non-standard amino acids into proteins could significantly expand the repertoire of designer enzymes."
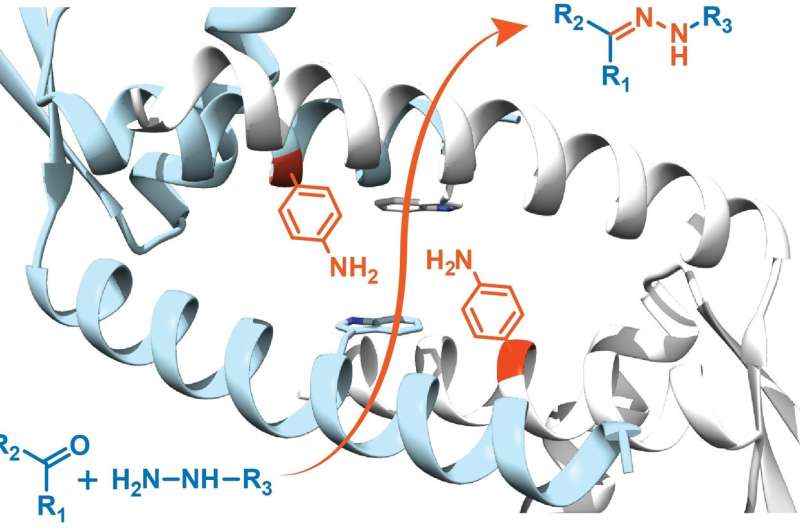
His team used a bacterial transcription factor as a starting point for their design. The LmrR protein is a transcriptional regulator in the bacterium Lactococcus lactis, which responds to antibiotics; it does not catalyse chemical reactions by itself. It is a doughnut-shaped dimer with a central opening that binds hydrophobic molecules. Using expanded genetic code techniques, Roelfes and his colleagues introduced the unnatural amino acid p-aminophenylalanine to the protein in such a way that the aniline of the side chain was present in the central opening. "Aniline is a known catalyst in chemical synthesis, but not a very potent one. Introducing it to the LmrR protein, increased catalysis by almost three orders of magnitude."
Different reactivities
The new enzyme links organic molecules by forming a hydrazone structure, a reaction which is used in medical biotechnology, for example, to couple drugs to antibodies. "In our experiments, we used a reaction that gives a nicely coloured product, so we could easily measure the activity of the new enzyme." Roelfes and his team are now modifying the central opening to increase the efficiency of the enzyme and tailor it to specific reactions.
"There are many options to use this enzyme design in chemical synthesis," says Roelfes "And we could also introduce other unnatural amino acids which have different reactivities." By combining these novel catalytic properties with the promiscuous binding site of the LmrR protein, Roelfes is confident that many new useful enzymes can be created.
More information: Ivana Drienovská et al, A designer enzyme for hydrazone and oxime formation featuring an unnatural catalytic aniline residue, Nature Chemistry (2018). DOI: 10.1038/s41557-018-0082-z
Journal information: Nature Chemistry
Provided by University of Groningen