Supercomputing simulations help reveal traffic mechanisms through the nuclear pore complex
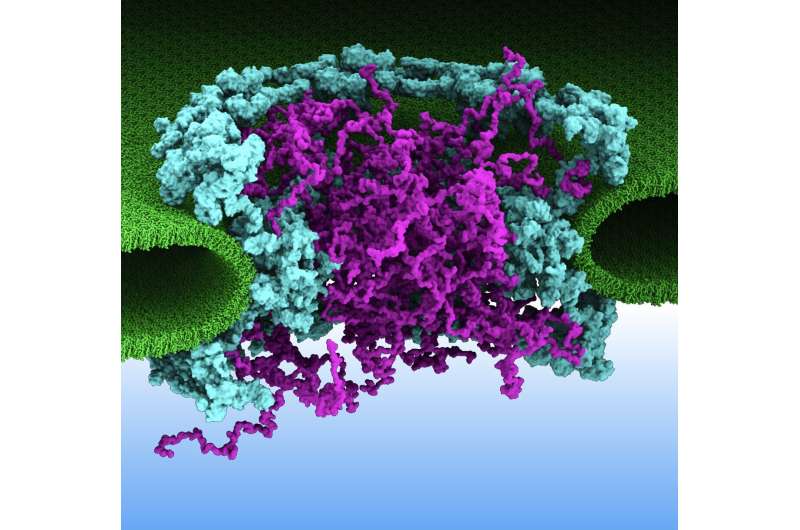
Students learn about the nucleus in ninth grade biology—it's the inner sanctum of biological cells, where the genome resides with the blueprints for cells to make proteins that are the building-blocks of life. Guarding those valuable blueprints is a dual-layer membrane, which not only protects, but also filters important molecules that regulate cellular functions.
Apertures called nuclear pore complexes (NPC) perforate the otherwise iron-clad membrane and act like crossing guards for macromolecular traffic in and out of the nucleus. If the crossing-guard misfires, it can cause human diseases such as cancer, viral infections, and neurodegenerative conditions.
A new mechanism has been determined for the first time for the passive transport of biomolecules through the nuclear pore complex. The work was published in the journal Nature Communications in August 2022.
The research team developed their NPC model through supercomputer simulations on the Frontera and Stampede2 systems of the Texas Advanced Computing Center (TACC)—and hope their work will help guide the development of future therapeutics.
"Our main finding is that a mesh-like interior of the nuclear pore exhibits a switch-over behavior based on protein size changing from a soft barrier for small proteins to a hard barrier beyond a certain threshold, essentially making it very difficult for proteins to get through," said study co-author David Winogradoff.
Winogradoff completed the study as a postdoctoral research associate working with co-author and professor Aleksei Aksimentiev in the Department of Physics at the University of Illinois at Urbana-Champaign. He's now a computational polymer chemist with the U.S. Food and Drug Administration.
Developing the model
Winogradoff's team used brute force simulations to study the kinetics of the nuclear pore transport at the time scale of tens of milliseconds, a phenomenal achievement for a system of nominally two hundred million atoms.
To do this, they combined several existing models, in particular the coarse-grained model developed by the Onck Group of the University of Groningen. The coarse-grained simulations resolved the disordered dynamics of only groups of atoms versus all-atom simulations, where every single atomic interaction is resolved.
The nuclear envelope model was also informed by cryogenic-electron microscopy data on two different scaffolding structures, a composite structure from Lynn2016 and the yeast structure of Kim2018.
"The model itself is in a way a separate accomplishment," said Aksimentiev.
"With the coarse-grained model, we saw spontaneous transport of proteins through this fluctuating mesh of filaments that extend into the cytoplasm. That is something that would be very, very difficult to probe using all-atom computational approaches," Aksimentiev added.
From those unbiased simulations the team then observed rare and rapid crossing events.
"We observed that there's a switch-over by protein size, from there always being a continuous path to it being very rare for there to be a continuous path connecting the top and the bottom of the nuclear pore through its central channel," Winogradoff said.
Supercomputer resources
The graphics processing unit (GPU) nodes on TACC's Frontera supercomputer ran software developed by the Aksimentiev Lab called Atomic Resolution Brownian Dynamics (ARBD).
Additionally, the scientists used TACC's Stampede2 and the Blue Waters system of the University of Illinois at Urbana-Champaign, both awarded through the National Science Foundation's Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS), formerly known as the Extreme Science and Engineering Discovery Environment (XSEDE).
"What really helped in having Frontera and Stampede2 resources available was that we were able to simulate a large range of sizes of proteins," Winogradoff said. "It sped up the process, which just took days versus months of/from local resources. Being able to run multiple replicas under different conditions gave a lot more weight to the robustness of our results," he added.
The simulations ran during TACC's Texascale Days, where select teams are awarded full use of the Frontera system, the flagship computer supported by the NSF and currently the top academic supercomputer at any university in the U.S.
Winogradoff's team scaled their NAMD simulations of the all-atom nuclear pore systems to about half of the nodes of Frontera, totaling about 250,000 processors.
Potential drug therapies and next steps
"In many ways I see this study as something that could provide some guidelines for future therapeutic development," Winogradoff said.
Cutting-edge emerging technologies aim to deliver drug therapies directly into the cell nucleus, which is difficult to reliably cross into.
"This research could provide information about the size threshold of drug cargo and whether, if it's too big, facilitate transport involving other proteins might be needed," Winogradoff said.
This work is a first step towards the long term goal of examining how the nucleus operates, according to Aksimentiev. His lab is working on an all-atom simulation that includes a variety of proteins that mimic a realistic, dense protein environment inside a living cell.
Aksimentiev concluded that "supercomputers are a unique tool that let you see what individual atoms are doing. From that, you can see how the behavior of individual atoms projects to the properties of larger scale molecular machines. This is something that we can presently do only by using supercomputers."
More information: David Winogradoff et al, Percolation transition prescribes protein size-specific barrier to passive transport through the nuclear pore complex, Nature Communications (2022). DOI: 10.1038/s41467-022-32857-1
Journal information: Nature Communications
Provided by University of Texas at Austin