Previously unknown mode of bacterial growth discovered
Bacteria as unicellular organisms reproduce by dividing their entire organism. In this way, they can multiply particularly quickly, which allows the exponential growth of bacterial populations, also known from pathogens. The population growth is based on the growth of the individual bacterial cells. It is also characterized by a rapid increase in size and, depending on the species, limited by a certain maximum cell size of the bacterium. Until now, research has assumed that a bacterial cell, just like its population as a whole, grows exponentially until it reaches this final size.
At Kiel University, the Microbial Biochemistry and Cell Biology research group led by Professor Marc Bramkamp investigates, among other things, bacterial organizational and reproductive mechanisms whose generally applicable principles are also important in the development of complex multicellular organisms. In a new research paper, Bramkamp and his team from the Institute of General Microbiology were able to show, using the bacterium Corynebacterium glutamicum as an example, that bacteria can also grow in a different, namely two-stage mode: Corynebacteria grow by synthesizing the cell wall at the cell poles. In this process, the growth rate of the cell poles is not identical at first after a cell division. The younger cell pole grows slower right after division. Due to this special model of elongation at the poles, cellular growth initially accelerates, and then transitions to steady, linear growth.
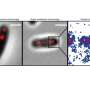
Based on special imaging techniques and a newly developed analytical method, the researchers succeeded in precisely quantifying the increase in size of the bacterial cells in the living organism. These data were incorporated into theoretical modeling, which revealed that the two-stage mode is the result of cell wall formation at the poles being the limiting process for growth in C. glutamicum. The Kiel researchers published the results today together with cooperation partners at the universities in Amsterdam and Munich in the scientific journal eLife.
Unusual bacterial growth observed and modeled
The regulation of growth and cell size is crucial for bacterial cell functions. Since it was previously assumed that single cells must increase their cell volume in proportion to their increasing protein content, growth was previously assumed to be predominantly exponential. This rapid increase in size goes hand in hand with the fact that bacteria must strictly control their cell size to guarantee morphologically uniform cells in a population. In fact, many bacterial species have complex mechanisms for regulating their size, but such mechanisms have not yet been identified in C. glutamicum and the closely related tuberculosis pathogen Mycobacterium tuberculosis.
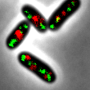
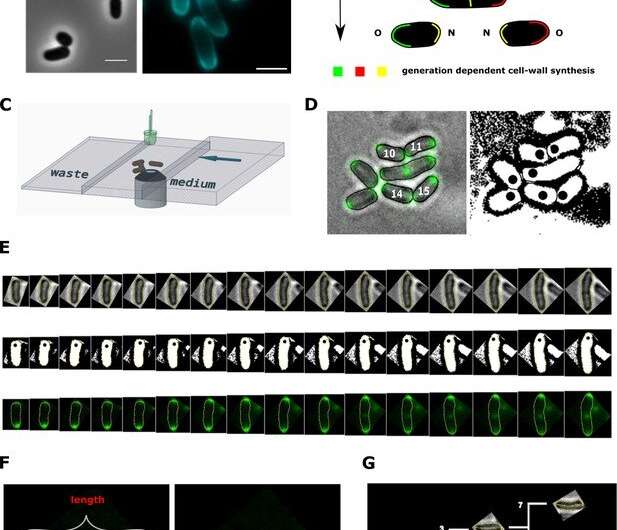
To investigate the specificity of this bacterial model organism, the Kiel research team observed its growth in vivo, i.e. on the living object. "An exciting aspect of Corynebacteria, which is already visible during microscopy, is that the cells do not always divide exactly symmetrically, as we know from many other bacteria," says Fabian Meyer, a doctoral student in Bramkamp's group. "Another special feature is that new cell wall does not develop on the long side of the cell, but is inserted exclusively at the cell poles," Meyer explains. Therefore, these bacteria are only elongated at the cell ends and do not increase their volume evenly over their entire length like many other bacteria.
To characterize this unusual growth behavior of C. glutamicum, the research team developed a theoretical model of the size development based on the Kiel data. With the help of biophysical methods, biophysicists Joris Messelink, and Professor Chase Broedersz from the Vrije Universiteit Amsterdam and the Ludwig-Maximilians-Universität München succeeded in extracting mean single-cell elongation patterns over time from this data, despite measurement noise and cell-to-cell variability.
"The calculations showed that the cells initially grow faster and faster, but then the increase in size slows down. In a way, there is a speed limit built into the growth dynamics of C. glutamicum," explains Broedersz. In contrast to the normally exponential course, the model here showed a two-step growth of the bacterial cells, which the researchers describe as asymptotically linear. "The limiting factor for the onset of linear growth is the type of cell wall formation that distinguishes C. glutamicum from other bacteria," Broedersz continues. The research team was then able to confirm this theoretical prediction using living bacteria: In bacteria in which a central but redundant growth enzyme was artificially switched off, slowing down the limiting factor for cell growth, the researchers repeated their growth inference procedure. The observed growth curves displayed an overall decrease of elongation rate while maintaining the two growth stages, in accordance with theoretical predictions.
Conclusions on bacterial evolution
"The modeling of the cell growth predicts precisely the results of the in vivo experiments, recapitulating the particularities of individual cell growth in C. glutamicum. In addition, the results offer an evolutionary interpretation of C. glutamicum's growth mechanisms. As for any organism, bacterial offspring must be similar to the parent generation in size and shape and bacteria have apparently evolved different growth strategies to achieve this aim," Bramkamp summarizes key findings of the interdisciplinary cooperation between cell biology and biophysics.
In the case of bacteria, this refers above all to cell length, the distribution of which should be relatively narrow in a population. "Our results show that the distribution of cell lengths is narrower in asymptotic linear growth than in exponential growth. The particular mode in C. glutamicum that transitions into linear growth could serve as a substitute for strict regulation of division length, since such linear growth is less sensitive to fluctuations. Therefore, it may not need its own growth control mechanism as we know it from other bacteria," Bramkamp continues.
More information: Joris JB Messelink et al, Single-cell growth inference of Corynebacterium glutamicum reveals asymptotically linear growth, eLife (2021). DOI: 10.7554/eLife.70106. elifesciences.org/articles/70106
Journal information: eLife
Provided by Kiel University