Researchers investigate previously unknown reproduction mechanism in Corynebacterium glutamicum
In recent decades, the interdisciplinary field of life sciences has made enormous progress, particularly in researching the molecular architecture of life. For example, extensive details of the genetic blueprint of many organisms are now known. Nevertheless, in areas such as cellular biology, many questions about fundamental life processes remain unanswered. One of these fundamental questions is how exactly individual cells, and thus ultimately also complex organisms, derive their spatial structures from biochemical information, and pass on the associated information to their offspring in a stable manner. At Kiel University (CAU), the Microbial Biochemistry and Cell Biology working group led by Professor Marc Bramkamp tackles this question, among other topics. In their latest research, Bramkamp and his team were able to gain new insights into how microorganisms reproduce their genetic information, and thereby organise their growth and reproduction, based on the example of the bacterium Corynebacterium glutamicum. Together with international colleagues, including from the Institut Pasteur in Paris, the Kiel research team published their new findings today in the renowned scientific journal Nature Communications.
How spatial structures arise from biochemistry
The growth of microorganisms is a relatively simple process, through which the question of reproduction and spatial organisation of organisms can be investigated. For example, many bacteria develop a characteristic bacillary (i.e. rod-shaped) form, like the bacterium Corynebacterium glutamicum investigated in the study. In order to better understand the underlying processes for inheritance and growth, the CAU researchers investigated how the chromosomes with the bacterial genes are compacted inside the cells and passed on to the offspring cells during the cellular cycle of the bacteria. Thus, they followed cells from birth right up to cell division. Bacteria reproduce via cell division, and must replicate their genetic information during this process. Overall, the associated processes in C. glutamicum and related species have hardly been investigated.
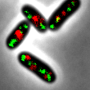
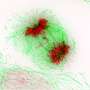
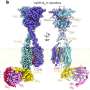
To distribute the replicated genetic information, the bacterium uses the so-called ParB protein (from "partitioning"). This protein binds the chromosomes to a specific DNA sequence, and links this to the cell poles, i.e. to both ends of the rod-shaped bacteria. In doing so, each chromosome is linked to a specific cell pole. Contrary to what was previously assumed for bacteria, Corynebacteria have two copies of each gene, just like most vertebrates including humans. With the help of ParB and a partner protein ParA, the duplicated chromosomes are then moved over the existing chromosomes. "The bacterial chromosome is therefore also important as a structural element for the organisation of the cell, and not only for the storage of genetic information," explained Bramkamp. "We were able to show that in this bacterium, the replication of the chromosomes essentially starts at the cell poles, and the new DNA is transported from there to the cell centre, and is thus divided particularly efficiently," continued Bramkamp.
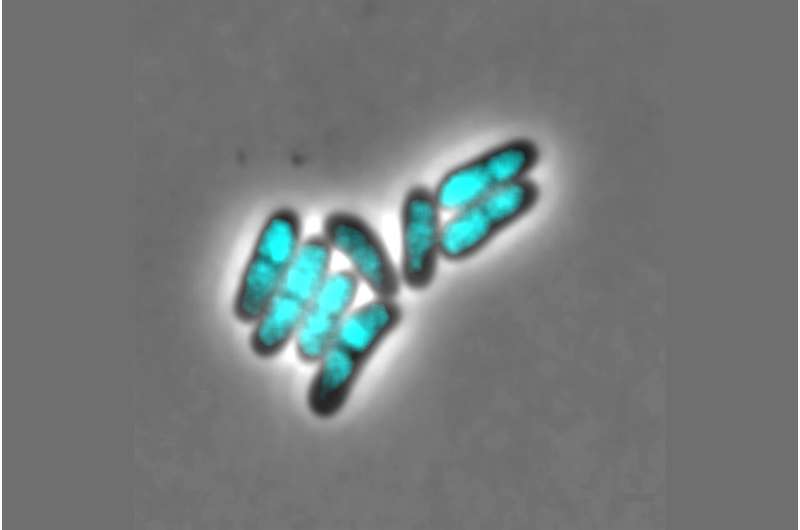
The chromosome is a strand of DNA which is many times larger than the size of a bacterial cell. In order to accommodate the genetic information in the cell, it must therefore be packaged and condensed. There are so-called condensin complexes involved in this process. These are specific enzyme complexes which assist with the organisation of the chromosomes and are added to the DNA by the ParB protein. The Kiel research team discovered that two different condensin complexes occur in C. glutamicum, but that both are not involved in the packaging of the genetic information. Instead, a newly-discovered condensin system regulates the proliferation of so-called plasmids in the bacterial cells. Plasmids are genetic elements which are present as ring-shaped DNA outside the chromosomes in the bacterial cell. Plasmids are important due to their involvement in so-called horizontal gene transfer, for example, such as the exchange of genetic information between different microbial species. This enables the spread of genes for antibiotic resistance and so-called virulence factors, i.e. characteristics which determine the pathogenic effect of a bacterium.
Making life processes visible in high resolution
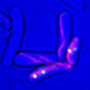
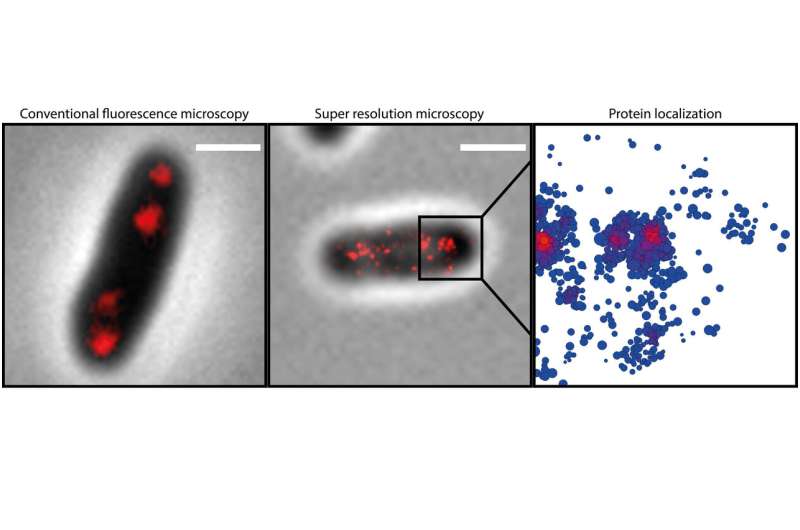
Bramkamp, who moved from the Ludwig-Maximilians-Universität München (LMU Munich) to the Institute of General Microbiology at the CAU last year, places particular emphasis on the development and use of innovative imaging techniques while researching the mechanisms of bacterial reproduction. While investigating the newly-discovered mechanisms, the research team used so-called single-molecule localisation microscopy (SMLM), which enables microscopic visualisation of single molecules with a precision of 20 nanometres. "In this way, we were able to make previously-unknown nanostructures visible which are involved in the organisation of the bacterial chromosome," emphasised doctoral researcher Giacomo Giacomelli. "The ability to observe and investigate the processes as they occur in the living organism opens up completely new perspectives for understanding the molecular mechanisms involved," continued Giacomelli.
Overall, the findings based on the example of C. glutamicum are of high scientific value, since this previously hardly-explored bacterium potentially represents a new model organism for the investigation of bacterial life processes. This offers highly-promising potential in both economic and medical terms. As such, in biotech production C. glutamicum plays an important role in manufacturing amino acids. It is also closely related to certain pathogens, like the tuberculosis pathogen Mycobacterium tuberculosis. Therefore, in both fields, an ever-growing understanding of the life processes, and in particular the reproduction of the bacterium, is of great importance.
More information: Kati Böhm et al. Chromosome organization by a conserved condensin-ParB system in the actinobacterium Corynebacterium glutamicum, Nature Communications (2020). DOI: 10.1038/s41467-020-15238-4
Journal information: Nature Communications
Provided by Kiel University