Researchers identify missing 'switch' that controls essential genes
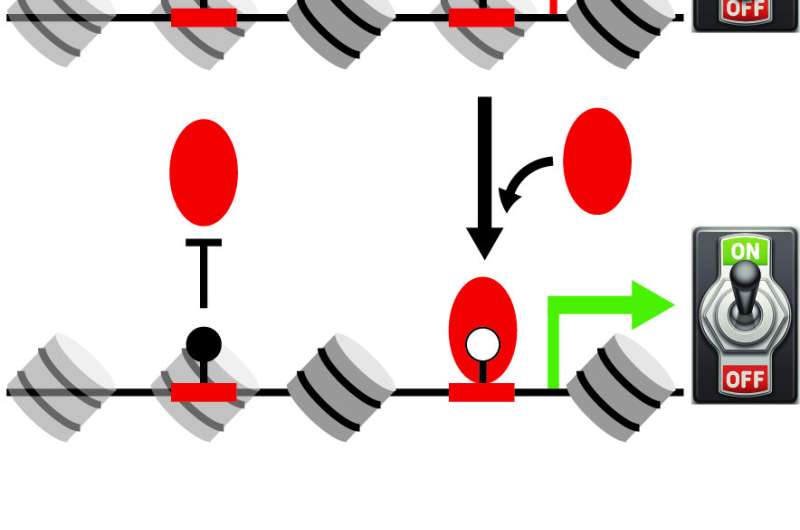
Proteins known as transcription factors act as switches that regulate the expression of nearby genes, but the identity of some of these genetic levers has so far remained mysterious. Now, researchers from the Schübeler group have pinpointed a new switch that regulates essential genes in the mouse and the human genome. Identifying missing gene switches and their function is critical to fully understand the molecular basis of health and disease.
If the human genome were a company, transcription factors would be the top-level managers, controlling when and how much genes are turned on in specific cells. These proteins typically bind short strings of DNA called 'motifs'. Scientists estimate that there are up to 2,800 transcription factors, but binding motifs have been identified for only about 800 of them.
One DNA motif that is bound by no known transcription factor is called the CGCG element, as it contains two cytosine nucleotides sitting next to guanine nucleotides. This motif is associated with highly expressed genes across human tissues and is commonly found within specific DNA regulatory regions where most of our genes start to be read.
However, discovering which transcription factors bind specific DNA sequences in living cells has been challenging, since regulatory regions usually contain several motifs. To look at proteins sitting on the CGCG motif, Ralph Grand and Lukas Burger, two researchers in the group of Dirk Schübeler, turned to a technique called single-molecule footprinting, which had been previously advanced by the Schübeler group. By mapping DNA regions that are blocked by proteins and those that are not, the technique allowed the team to discover a 'footprint' of an unknown factor bound to the CGCG motif.
To identify the factor associated with this footprint, the researchers opened up the nuclei of living cells and pulled out their innards. Then, they used the CGCG motif as a bait to fish out the proteins bound to it. Using mass spectrometry, a technique that identifies molecules by their mass and charge, the researchers detected the Btg3-associated nuclear protein (BANP) as the only protein bound to the CGCG motif.
"This protein has been known before, but it was thought to repress gene activity at the periphery of the nucleus," Grand says. "We show that it does quite the opposite: it's a very potent activator of gene expression."
Hiding in plain sight
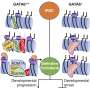
The team found that BANP has a high affinity for the CGCG motif, both in mouse and human cells. Removing BANP in stem cells causes a decrease in the expression of several genes, including essential ones involved in key biological processes such as transcription, DNA replication, and the assembly of chromatin—the complex of DNA and proteins that forms chromosomes. The researchers observed similar drops in gene expression also when BANP was removed in neurons.
After binding to specific regulatory regions within the genome, BANP makes the DNA accessible to other proteins. This likely helps regulatory factors to bind and control gene expression. The findings, published today in Nature, could redefine how essential genes are controlled. "These genes, which are expressed in every cell of the body but at different levels, could be regulated by the same switch present in all cells rather than by a series of transcription factors across different cell types", Grand says.
Despite its key role in regulating gene expression, BANP had been hiding in plain sight. "We believe that is because BANP is so essential: you touch it, and the cell dies," Schübeler says. "This made it hard to identify it by any kind of genetic screening approach, which makes us wonder whether there are more of these factors out there that are invisible to us for the same reasons," he adds.
Cancer link
Further experiments showed that BANP binds DNA only when the CGCG motif is not methylated. DNA methylation is a chemical modification that can repress gene activity. In human cancer cells, which show abnormal patterns of DNA methylation, BANP binds to regulatory regions containing unmethylated CGCG motifs but not to those containing methylated motifs. "That opens up an interesting idea," Burger says. "DNA methylation might regulate where BANP can bind, thus influencing how genes are expressed," he adds.
Understanding how BANP and other factors bind DNA could have important implications in biomedicine, as genetic variation in regulatory regions can determine whether some individuals are more susceptible to disease, Schübeler says. The more switches scientists characterize, the better they'll understand what type of information is contained in regulatory regions of the genome, how altering this information can result in disease, and how these switches can be used to control genes.
The study was done in collaboration with the Thomä lab and the Proteomics and Genomics technology platforms at the FMI, as well as the Vermeulen lab at the Radboud Institute for Molecular Life Sciences in Nijmegen, the Netherlands.
More information: Ralph S. Grand et al, BANP opens chromatin and activates CpG-island-regulated genes, Nature (2021). DOI: 10.1038/s41586-021-03689-8
Journal information: Nature