Study reveals dynamics of crucial immune system proteins
Of the many marvels of the human immune system, the processing of antigens by the class I proteins of the major histocompatability complex (MHC-I) is among the most mind-boggling. Exactly how these proteins carry out their crucial functions has not been well understood. Now, however, researchers at UC Santa Cruz have worked out the details of key molecular interactions involved in the selection and processing of antigens by MHC-I proteins.
The new findings, published December 3 in Proceedings of the National Academy of Sciences, help explain certain puzzling differences among MHC-I proteins, with implications for understanding autoimmune diseases and immune responses to infections and cancer. The results also suggest ways in which MHC-I proteins can be manipulated in the laboratory for use in diagnostic and therapeutic applications.
"Our discovery of these fundamental mechanisms enables us to develop technologies with tremendous potential for diagnostic and therapeutic purposes," said Nikolaos Sgourakis, assistant professor of chemistry and biochemistry at UC Santa Cruz and corresponding author of the paper.
The role of MHC-I proteins is to enable every cell in the body to display on its surface fragments of all the proteins being produced in that cell, typically about 10,000 different proteins. The protein fragments displayed by MHC-I proteins on the cell surface are scanned by specialized immune cells called cytotoxic T cells, which can recognize foreign proteins from an infection or mutated proteins from a tumor and launch an immune response.
"The cell has this barcoding system in place so it can show the immune system what's going on inside, and the T cells continuously surveil the surfaces of cells to sniff out the barcodes of aberrant proteins," Sgourakis explained.
Sgourakis and his team, working in close collaboration with coauthor Erik Procko's group at the University of Illinois, focused on the process of "antigen loading"—how the protein barcodes are selected and bound to MHC-I proteins so they can be displayed on the cell surface. Molecular "chaperones" play an important role in antigen loading and help determine which protein fragments get displayed. The new paper reveals how the interplay between MHC-I proteins and chaperones shapes the repertoire of displayed antigens.
There are thousands of different variants of the human MHC-I proteins, produced by different "alleles" of the MHC-I genes. The extreme variability of MHC-I proteins accounts for much of the individual variation in immune responses, including differences in susceptibility to autoimmune diseases, infections, and cancer. Each person has six main MHC-I alleles (three inherited from mom and three from dad), and each allele can display a unique subset of all possible barcodes.
"Our six MHC-I proteins sample a fraction of all the possible barcodes being generated in our cells. The ones they select become the displayed antigen repertoire, which is different for every person," Sgourakis said.
Sgourakis's team studied four different MHC-I alleles, examining their interactions with molecular chaperones and antigens. One function of the chaperones is to help MHC-I proteins fold into their active shapes and stabilize them to prevent misfolding and aggregation. But only some MHC-I alleles are dependent on chaperones for antigen loading. The new findings explain why that is and reveal important details of the antigen selection process.
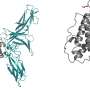
The key to understanding these interactions was the use of nuclear magnetic resonance (NMR) techniques to reveal dynamic structural changes in the MHC-I proteins. "We've had static crystal structures of MHC proteins, but we could not figure out why some are chaperone-dependent and others are not," Sgourakis said. "It turns out to be a matter of protein dynamics."
The researchers found that if the three-dimensional structure of the MHC-I molecule is rigid, chaperones are not involved in antigen loading. If it has flexibility in the peptide binding groove, however, the chaperone will interact with it and help with the antigen loading process. The chaperone can eject antigens that have low affinity for the binding groove, ensuring that the MHC-I protein binds only high-affinity antigens that can be displayed at the cell surface in the proper conformation to activate a T cell response.
A flexible groove may enable the MHC-I molecule to accommodate a broader range of antigens, Sgourakis said. "The immune system has to cover all these possible barcodes with a limited number of MHC-I alleles. One way to do that is for the binding groove to adopt different shapes, but that flexibility comes at a price. You need to have a mechanism to stabilize these more flexible proteins—hence the chaperones," he said.
Sgourakis said his lab can now use chaperones in a high-throughput procedure to create libraries of barcoded MHC-I protein complexes encompassing hundreds of different peptides for use in screening T cells from patients and determining their antigen specificities. This procedure has potential applications in immunotherapy for cancer and other diseases. Sgourakis said his team is actively exploring this direction for cancer immunotherapy development in collaboration with clinical researchers at Children's Hospital of Philadelphia.
More information: Molecular determinants of chaperone interactions on MHC-I for folding and antigen repertoire selection, Proceedings of the National Academy of Sciences (2019). DOI: 10.1073/pnas.1915562116 , www.pnas.org/content/early/2019/12/02/1915562116
Journal information: Proceedings of the National Academy of Sciences
Provided by University of California - Santa Cruz