Biochemists discover mechanism that helps flu viruses evolve

Influenza viruses mutate rapidly, which is why flu vaccines have to be redesigned every year. A new study from MIT sheds light on just how these viruses evolve so quickly, and offers a potential way to slow them down.
The MIT team found that flu viruses' rapid evolution relies in part on their ability to hijack some of the cellular machinery of the infected host cell—specifically, a group of proteins called chaperones, which help other proteins fold into the correct shape. When the viruses were unable to get help from these chaperones, they did not evolve as rapidly as when they could obtain extensive help from host chaperones. Moreover, the specific evolutionary trajectories followed by individual flu proteins depend on host chaperone activities.
The findings suggest that interfering with host cell chaperones could help prevent flu viruses from becoming resistant to existing drugs and vaccines, says Matthew Shoulders, the Whitehead Career Development Associate Professor of Chemistry at MIT.
"It's relatively easy to make a drug that kills a virus, or an antibody that stops a virus from propagating, but it's very hard to make one that the virus doesn't promptly escape from once you start using it," Shoulders says. "Our data suggest that, at some point in the future, targeting host chaperones might restrict the ability of a virus to evolve and allow us to kill viruses before they become drug resistant."
Shoulders is the senior author of the study, which is a collaborative effort with Leonid Mirny, a professor of physics at MIT; and Yu-Shan Lin, a professor at Tufts University. Angela Phillips, an MIT graduate student and National Science Foundation graduate fellow, is the lead author of the paper, which appears in the journal eLife on Sept. 26.
A little help
Flu viruses carry eight genome segments, all encoded by RNA. Of particular interest to flu researchers is the gene for the hemagglutinin protein, which is displayed on the surface of the viral envelope and interacts with cells of the infected host. Most flu vaccines target this protein, but these vaccines have to be updated every year to keep up with the protein's ability to evolve quickly.
However, this rapid evolution also poses a challenge for the viruses themselves. When proteins mutate, they may become unable to fold into the shape they need to assume to perform their function. Previous research, such as the pioneering work of the late Susan Lindquist, a biology professor at MIT, has shown that in many organisms, evolution of endogenous proteins depends on the ability of that organism's chaperones to help mutated proteins fold.
In this study, the MIT team investigated whether viruses can take advantage of their host's chaperone proteins to help with their own evolution.
"Viral proteins are known to interact with host chaperones, so we suspected that this interplay could have a major impact on what evolutionary pathways are available to the virus," Shoulders says.
To test their hypothesis, the researchers generated one set of cells with low protein-folding activity by inhibiting a key chaperone protein called heat shock protein 90 (Hsp90). In another set of cells, they used chemical genetic methods previously developed by Shoulders to enhance the levels of numerous chaperone proteins, creating a cellular environment with high protein-folding activity.
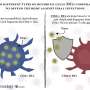
The researchers infected both sets of cells, plus a group of cells with normal chaperone levels, with a strain of flu and then allowed the virus to evolve for nearly 200 generations. They found that the virus did indeed evolve faster in the cells with higher chaperone levels than in the cells with inhibited chaperone proteins.
"This finding suggests that influenza will acquire new traits that might be beneficial for it faster when you have the heat shock response activated, and slower when you have key chaperones inhibited," Shoulders says.
Blocking escape routes
The researchers also identified specific proteins that tend to become more mutated in cells with more chaperones. One of these is the hemagglutinin protein, and another is an enzyme called PA, which is a type of RNA polymerase that helps the virus copy its genes. The team also identified specific amino acids within these proteins that are more likely to become mutated in different protein-folding environments.
"The authors develop very nice chemical genetic tools for precisely manipulating proteostasis in human cells, and the application of their methods led to a number of interesting findings," says Jesse Bloom, a viral evolution expert and associate member of the Fred Hutchinson Cancer Research Center, who was not involved in the research. "Perhaps the most compelling is the identification of a specific mutation in influenza (H452Q in PA) that has different effects depending on whether the heat shock response is activated versus whether Hsp90 is inhibited. Identification of this mutation is proof of principle that a virus' ability to tolerate specific mutations can be affected by chaperones, providing the first link between host proteostasis and viral evolution."
Targeting this phenomenon could offer a way to delay viral evolution and decelerate escape from existing drugs and vaccines, the researchers say. Many chaperone inhibitors already exist, and some are now being tested in clinical trials to treat cancer and some viral infections. The new data imply that treating patients with a chaperone-inhibiting drug along with another antiviral therapy, such as a drug or vaccine, could help ensure that the virus does not evolve resistance to the therapeutic.
The researchers believe this phenomenon is likely also found in other viruses, and they are now studying HIV, another virus that mutates rapidly. They also plan to study how a host cell's protein-folding capacity may affect the evolution of antiviral drug or antibody resistance, using therapeutics that circulating viruses are already resistant to.
"We can recapitulate environmental pressures like antiviral drugs in the lab, in the context of different host protein-folding environments, and see whether there's a big impact. Our data suggest that there's going to be, but we have to actually test it out," Shoulders says.
Journal information: eLife
Provided by Massachusetts Institute of Technology
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.