Researchers characterize Ssb chaperone protein for the first time

For 20 years, researchers have tried to find out why the chaperone Ssb is the only member of the widespread Hsp70-chaperone family that is able to bind directly to the ribosome. This question has now been answered at the University of Konstanz. The researchers identified two regions within the Ssb chaperone that mediate direct contact with the ribosome and thus support the function of Ssb. "We can demonstrate that Ssb interacts with the ribosome mainly via positively charged amino acids", says Professor Elke Deuerling, whose team discovered the binding sites of Ssb. The results were published on 5 December 2016 in Nature Communications.
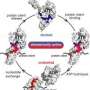
The chaperone Ssb is a member of the highly conserved Hsp70 chaperone family found in all living organisms. Hsp70 chaperones play a central role for preventing abnormal protein folding as well as for the transport of proteins. Ssb is the only family member that is in direct contact with the ribosome. It is active at a very early stage, when new proteins are being synthesised in the ribosome. "This ability is unique and cannot be detected at first glance when comparing Ssb with other Hsp70 chaperones. Only a few amino acids provide Ssb with this additional feature and show how flexible Hsp70 chaperones can be," explains molecular biologist Elke Deuerling. Dr. Anne Hanebuth significantly contributed to the discovery of the Ssb attachment sites in her doctoral thesis.
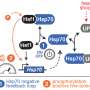
In several experiments, the biologists found out that the Ssb attachment sites are not essential under normal laboratory conditions. If they are mutated, nothing changes for the yeast cells. However, if co-chaperone RAC is absent, abnormal protein folding occurs in Ssb mutant cells, resulting in pronounced cellular defects. "We believe that these multivalent interactions with direct attachment sites and the cooperation with RAC allow the chaperone Ssb to position itself at the ribosome in an optimal way," says Deuerling. Finding this exact position at the ribosome is of crucial importance for the effectiveness of the chaperone.
In contrast to the co-chaperone RAC, the ribosome-bound chaperone Ssb can only be found in fungi such as yeast. In higher cells, RAC probably cooperates with other Hsp70 chaperones. This is why the researchers assume that the RAC-Hsp70-chaperone system is generally very important for correct folding of proteins and for keeping the cells fit. "How the RAC-Hsp70 system operates in higher cells and what effects it has on proteins relevant for diseases will be a huge topic in the SFB," says Deuerling. She further comments on the Ssb-chaperone results: "It was great team work within the University of Konstanz and with renowned teams from Heidelberg and Stanford. Computer simulations of the molecular dynamics of Ssb provided important indications of molecular interactions within the Ssb protein. This led to hypotheses about the ribosome interaction, which we have studied using further genetic, biochemical and kinetic approaches. Step by step, we were able to achieve our results."
More information: Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction. Nature Communications DOI: 10.1038/ncomms13695
Journal information: Nature Communications
Provided by University of Konstanz