This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Examining the role of 'blind dating' in bacteria evolution
Proteins are the key players for virtually all molecular processes within the cell. To fulfill their diverse functions, they have to interact with other proteins. Such protein-protein interactions are mediated by highly complementary surfaces, which typically involve many amino acids that are positioned precisely to produce a tight, specific fit between two proteins. However, comparatively little is known about how such interactions are created during evolution.
Classical evolutionary theory suggests that any new biological feature involving many components (like the amino acids that enable an interaction between proteins) evolves in a stepwise manner. According to this concept, each tiny functional improvement is driven by the power of natural selection because there is some benefit associated with the feature. However, whether protein-protein interactions also always follow this trajectory was not entirely known.
Using a highly interdisciplinary approach, an international team led by Max Planck researcher Georg Hochberg from the Terrestrial Microbiology in Marburg have now shed new light on this question. Their study provides definitive evidence that highly complementary and biologically relevant protein-protein interactions can evolve entirely by chance.
Proteins cooperate in a photoprotection system
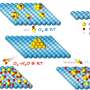
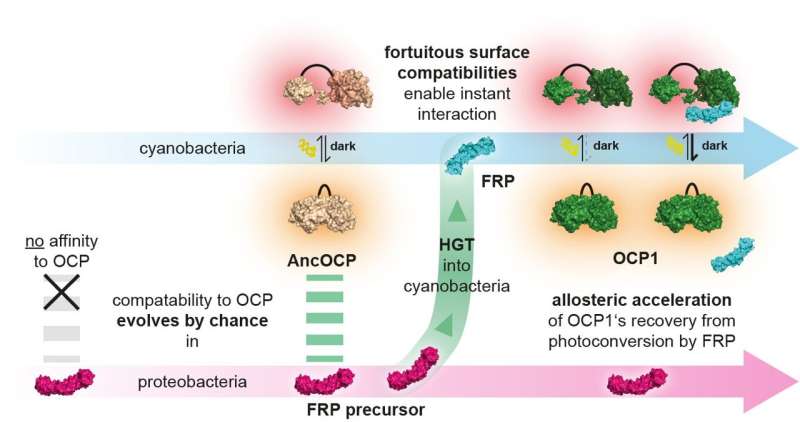
The research team made their discovery in a biochemical system that microbes use to adapt to stressful light conditions. Cyanobacteria use sunlight to produce their own food through photosynthesis. Since much light damages the cell, cyanobacteria have evolved a mechanism known as photoprotection: if light intensities become dangerously high, a light intensity sensor named Orange Carotenoid Protein (OCP) changes its shape.
In this activated form, OCP protects the cell by converting excess light energy into harmless heat. In order to return into its original state, some OCPs depend on a second protein: The Fluorescence Recovery Protein (FRP) binds to activated OCP1 and strongly accelerates its recovery.
"Our question was: Is it possible that the surfaces that allow these two proteins to form a complex evolved entirely by accident, rather than through direct natural selection?" says Georg Hochberg.
"The difficulty is that the end result of both processes looks the same, so we usually cannot tell why the amino acids required for some interaction evolved—through natural selection for the interaction or by chance. To tell them apart, we would need a time machine to witness the exact moment in history these mutations occurred," Hochberg explains.
Luckily, recent breakthroughs in molecular and computational biology has equipped Georg Hochberg and his team with a laboratory kind of time machine: ancestral sequence reconstruction.
In addition, the light protection system of cyanobacteria, which is under study in the group of Thomas Friedrich from Technische Universität Berlin since many years, is ideal for studying the evolutionary encounter of two protein components. Early cyanobacteria acquired the FRP proteins from a proteobacterium by horizontal gene transfer. The latter had no photosynthetic capacity itself and did not possess the OCP protein.
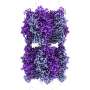
To work out how the interaction between OCP1 and FRP evolved, graduate student Niklas Steube inferred the sequences of ancient OCPs and FRPs that existed billions of years ago in the past, and then resurrected these in the laboratory. After translation of the amino acid sequences into DNA he produced them using E. coli bacterial cells in order to be able to study their molecular properties.
A fortunate coincidence
The Berlin team then tested whether ancient molecules could form an interaction. This way the scientists could retrace how both protein partners got to know each other. "Surprisingly, the FRP from the proteobacteria already matched the ancestral OCP of the cyanobacteria, before gene transfer had even taken place. The mutual compatibility of FRP and OCP has thus evolved completely independently of each other in different species," says Thomas Friedrich.
This allowed the team to prove that their ability to interact must have been a happy accident: selection could not plausibly have shaped the two proteins' surfaces to enable an interaction if they had never met each other. This finally proved that such interactions can evolve entirely without direct selective pressure.
"This may seem like an extraordinary coincidence," Niklas Steube says. "Imagine an alien spaceship landed on earth and we found that it contained plug-shaped objects that perfectly fit into human-made sockets. But despite the perceived improbability, such coincidences could be relatively common. But in fact, proteins often encounter a large number of new potential interaction partners when localisation or expression patterns change within the cell, or when new proteins enter the cell through horizontal gene transfer."
Georg Hochberg adds, "Even if only a small fraction of such encounters ends up being productive, fortuitous compatibility may be the basis of a significant fraction of all interactions we see inside cells today. Thus, as in human partnerships, a good evolutionary match could be the result of a chance meeting of two already compatible partners."
The work is published in the journal Nature Ecology & Evolution.
More information: Niklas Steube et al, Fortuitously compatible protein surfaces primed allosteric control in cyanobacterial photoprotection, Nature Ecology & Evolution (2023). DOI: 10.1038/s41559-023-02018-8
Journal information: Nature Ecology & Evolution
Provided by Max Planck Society