Surface protein editing in bacteria
University of Minnesota researchers have discovered this previously unknown signaling pathway that regulates surface proteins on bacteria that can lead to new targets for antibiotics.
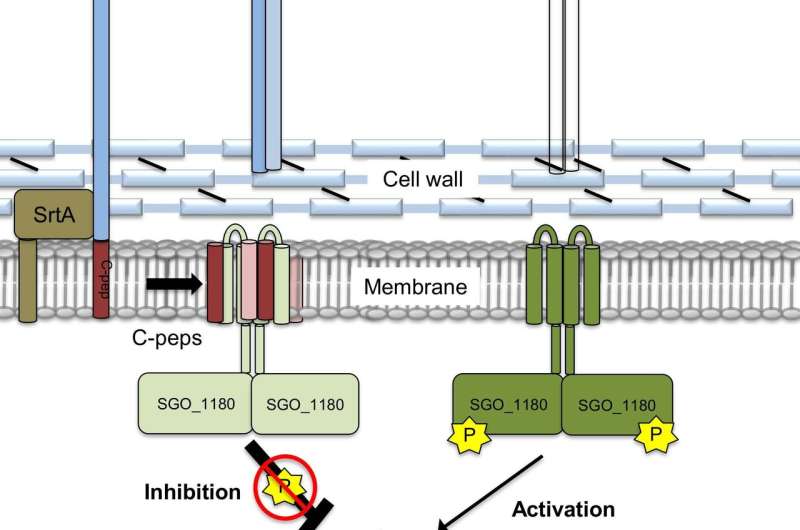
Researchers studied how oral bacteria adhere to and develop biofilms (plaque) in the oral cavity. The team wanted to learn whether and how the bacterial cells might adjust their adhesive surface proteins. They discovered a previously unknown circuit that is embedded in the cell membrane that can signal for changes in the surface adhesive proteins. This circuit appears to be conserved among a subset of Gram-positive bacteria. The intramembrane bacterial signaling system calls for different surface proteins to compensate in the absence of primary surface proteins. This mechanism provides compensatory biofilm (plaque) formation.
This mechanism appears to function in microbial communities in vitro and in the human mouth. Genes encoding surface adhesive proteins differ when the bacteria are recovered from saliva versus dental plaque in the same person at the same time.
When activated, this circuit rescued biofilm formation—which is when microorganisms strongly attach and grow on a surface—helping bacteria to survive in dental plaque.
The results, published in Science Signaling, found:
- a previously unrecognized signaling system within the cell membrane that regulates surface adhesive protein gene expression;
- the signaling system calls for the regulation of different surface proteins as the available pool of surface proteins is altered;
- the regulatory signal is a conserved amino acid sequence found in fragments cleaved from the surface adhesive proteins. When the fragment is present in the membrane, the system is "off" and, when it is absent, the system is "on" and alternative surface proteins are expressed;
- this intramembrane signaling system appears to compensate as a "fail-safe" mechanism to edit surface proteins and enable the bacteria to adhere and colonize different body surfaces.
"Discovering this previously unknown signaling pathway that regulates surface proteins on bacteria may help us to understand better how complex microbial communities develop and offer new targets for antibiotics," said Dr. Mark Herzberg, a professor in the School of Dentistry and a member of the Masonic Cancer Center.
More information: An intramembrane sensory circuit monitors sortase A–mediated processing of streptococcal adhesins, Sci. Signal. 07 May 2019: Vol. 12, Issue 580, eaas9941, DOI: 10.1126/scisignal.aas9941 , stke.sciencemag.org/content/12/580/eaas9941
Journal information: Science Signaling
Provided by University of Minnesota