Broken mitochondria use 'eat me' proteins to summon their executioners
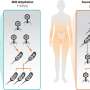
When mitochondria become damaged, they avoid causing further problems by signaling cellular proteins to degrade them. In a paper publishing April 11, 2019, in the journal Developmental Cell, scientists in Norway report that they have discovered how the cells trigger this process, which is called mitophagy. In cells with broken mitochondria, two proteins—NIPSNAP 1 and NIPSNAP 2—accumulate on the mitochondrial surface, functioning as "eat me" signals, recruiting the cellular machinery that will destroy them.
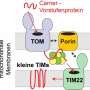
NIPSNAP 1 and 2 are normally found inside healthy mitochondria, although their function inside the cell is unknown. "When a cell's respiration chain is disrupted, and the mitochondria are damaged, import of these proteins into the matrix and inner membrane space of the mitochondria is interrupted," says senior author Anne Simonsen, a professor at the Department of Molecular Medicine at the Institute of Basic Medical Sciences of the University of Oslo. "In that case, the import system does not function and they remain bound to the surface of the damaged mitochondria signaling for mitophagy."
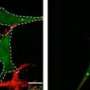
In this study, the researchers studied human HeLa cells where both NIPSNAP1 and NIPSNAP 2 function were eliminated. "When we do that, these cells cannot clear the mitochondria after damage," says Simonsen. However, in cells with functional NIPSNAP proteins, when mitophagy was induced through the addition of a chemical disruptor, they observed that the NIPSNAP proteins act in concert with the PINK and PARKIN proteins, proteins already known to have a role in triggering autophagy and to have a role in Parkinson's Disease.
PARKIN labels cells with ubiquitin, a small protein that directs the cells towards degradation. "Ubiquitin is the classical signal to recruit autophagy," says co-author Terje Johansen, of the University of Tromsø - The Arctic University of Norway. "What we saw is that in addition to ubiquitin, NIPSNAP proteins are required to recruit autophagy proteins; they are not targeted to the mitochondria unless these NIPSNAP proteins are found on the surface."
The team showed this finding has important physiological implications in vivo by investigating the NIPSNAP/PINK/PARKIN mechanism in a zebrafish animal model. They compared wild-type zebrafish and a fish line with reduced NIPSNAP1 protein abundance.
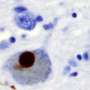
"We see that the mutant fish lacking adequate functional NIPSNAP1 are not able to move as the wild-type fish," says Simonsen. They have a Parkinsonian-like phenotype with reduced numbers of dopaminergic neurons. However, they could rescue this locomotion defect by adding L-dopa, the same compound used to treat human Parkinson's Disease, to the water.
Even more dramatically, animals entirely lacking NIPSNAP1 protein died within five days. "Clearly, clearance of mitochondria is important for the health of these dopaminergic neurons. That is particularly important since neurons generally cannot divide," says Johansen.
As evolutionarily conserved proteins, NIPSNAP proteins are found throughout the animal kingdom, including humans.
More information: Developmental Cell, Abudu, Pankiv, and Mathai et al.: "NIPSNAP1 and NIPSNAP2 act as "eat me" signals for mitophagy" www.cell.com/developmental-cel … 1534-5807(19)30224-2 , DOI: 10.1016/j.devcel.2019.03.013
Journal information: Developmental Cell
Provided by Cell Press