Researchers push the boundaries of optical microscopy

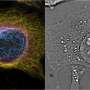
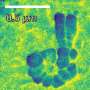
The field of optical microscopy research has developed rapidly in recent years. Thanks to the invention of a technique called super-resolution fluorescence microscopy, it has recently become possible to view even the smaller parts of a living cell. Now, by making a smart refinement to that technique, researchers at TU Delft have pushed its boundaries even further. Where previously objects measuring up to 10-20 nanometres could be observed, their method makes it possible to focus on structures of as tiny as 3 nanometres across.
Delft cloth trader and scientist Antoni van Leeuwenhoek's home-made microscopes had a resolution of less than a micrometre, which enabled him to observe structures like bacteria and sperm cells. But even in the seventeenth century, Van Leeuwenhoek was already approaching the so-called 'diffraction limit', a theoretical boundary beyond which two adjacent points cannot be distinguished under an optical microscope. This limit is determined in part by the wavelength of the light being used. According to the theory, the maximum size of the object you can image using a conventional microscope is half that wavelength. Anything smaller is impossible to bring into sharp focus.
The diffraction limit was long thought to a hard boundary, determined by the laws of nature. But by applying clever tricks, physicists eventually succeeded in crossing it. Not so long ago, in 2014, the Nobel Prize for Chemistry was awarded to the three researchers who invented the workaround, known as 'super-resolution fluorescence microscopy'. In this technique, certain proteins or molecules are made fluorescent by genetic modification. The weak light signal they emit can then be captured with the help of an optical microscope. "In practice, though," says researcher Bernd Rieger, "the problem with making proteins fluorescent is that you can't label all those of a particular type. Only 30-50 per cent of them, at most. When you then start taking measurements, you see only a number of individual luminous points, not the complete structure you are trying to view."
To solve the problem, the Delft researchers have devised an adaptation to super-resolution microscopy. This is comparable with what is known in photography as 'compositing': stacking multiple images to create a single compound picture. "Averaging the information from different measurements was already being done in electron microscopy," explains researcher Sjoerd Stallinga. "But that's a completely different technology. It took our doctoral candidate Hamidreza Heydarian two years to convert the technique for use in optical microscopy."
One problem was that combining hundreds, if not thousands, of 'snapshots' requires huge amounts of processing power. With a normal computer, it took several days to construct a clear image from all the data. "Fortunately," says Rieger, "thanks to the computer games industry, we have access to graphics cards able to calculate extremely well in parallel." A programmer from the Netherlands eScience Center in Amsterdam joined the project and converted an existing algorithm for normal PCs into one the researchers could run on such a graphics card. As a result, the measurements can now be combined into a single image within a few hours.
This research is narrowing the gap between electron and optical microscopy, which is important because the two techniques deliver different results and so are complementary, but are still a long way apart in terms of their possibilities. "The best electron microscopes are 30 to 50 times more powerful than the best optical ones," says Stallinga. "Bringing the two worlds closer together could lead to new biological insights."
According to the researchers, their technique – which is already achieving resolutions at the three-nanometre level – should eventually make it possible to view structures measuring just one nanometre. Below that threshold, the dimensions of the fluorescent labels become a limiting factor.
The findings have been published in the journal Nature Methods.
More information: Hamidreza Heydarian et al. Template-free 2D particle fusion in localization microscopy, Nature Methods (2018). DOI: 10.1038/s41592-018-0136-6
Journal information: Nature Methods
Provided by Delft University of Technology