Researchers successfully sequence total RNA of single cells
By combining a number of methods, researchers from the RIKEN Advanced Center for Computing and Communications (ACCC) in Japan have developed a method that allows full-length sequencing of the total RNA of a single cell. The ability to do such full-length sequencing is important for understanding how single cells develop and function in biological systems.
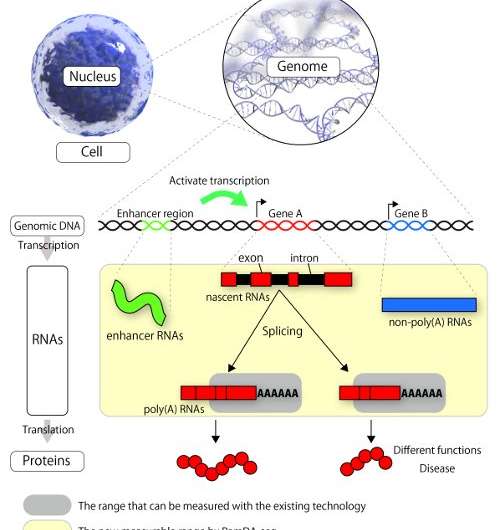
Great progress has been made in decoding the genome and transcriptome—RNA expression—in different species and organisms, but much work remains to be done to understand what is happening in individual cells. One of the key challenges up to now in single-cell RNA sequencing, a technology for decoding the transcriptome of a single cell, has been the difficulty in determining sequences of RNAs that are not polyadenylated—a type of modification that is found in almost all messenger RNAs. Non-polyadenylated RNAs have low expression and have slightly different structures. Recent studies have shown that non-polyadenylated RNAs are involved in diverse cell functions and diseases. Despite their importance, methods up to now have either been unable to detect the non-polyadenylated RNAs or only able to detect them with very low sensitivity.
To overcome this limitation, the RIKEN scientists combined a variety of methods to develop a system titled random displacement amplification sequencing (RamDA-seq), which is highly sensitive to both polyadenylated and non-polyadenylated RNAs and strongly immune to types of biases that often occur during sequencing. The results are published in Nature Communications.
Through a number of tests, they determined that RamDA-seq is very sensitive and also has good reproducibility. They found that it can provide full-length sequences of extremely long transcripts, and is very sensitive, unlike previous methods, to non-polyadenylated RNAs. They applied RamDA-seq to mouse embryonic stem cells and properly detected the sequence of known RNAs, as well as additional sequences that were not previously known.
Using RamDA-seq, the scientists discovered a number of interest facts. First, they showed that non-polyadenylated RNAs are dynamically regulated in a manner that depends on the differentiation state, so that a single cell in a different stage might express them differently. They also found a unique phenomenon—called recursive splicing, which has garnered attention in recent years—in certain genes, and were able to analyze these at the single-cell level. Finally, they found that RamDA-seq is useful for understanding the activity of enhancer RNAs—RNAs which reflect the functions of regulatory genomic regions.
According to Itoshi Nikaido, leader of the Bioinformatics Research Unit of ACCC, "We are very excited at the positive results we got from this new system, and very much hope that researchers around the world will be able to use it to give us new insights into gene expression in individual cells, allowing us to better understand the complexity of biological systems."
More information: Tetsutaro Hayashi et al. Single-cell full-length total RNA sequencing uncovers dynamics of recursive splicing and enhancer RNAs, Nature Communications (2018). DOI: 10.1038/s41467-018-02866-0
Journal information: Nature Communications
Provided by RIKEN