It takes two to tangle: Scientists further unravel telomere biology
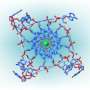
Chromosomes - long, linear DNA molecules – are capped at their ends with special DNA structures called telomeres and an assortment of proteins, which together act as a protective sheath. Telomeres are maintained through the interactions between an enzyme, telomerase, and several accessory proteins. Researchers at The Wistar Institute have defined the structure of one of these critical proteins in yeast.
Understanding how telomeres keep chromosomes – and by extension, genomes – intact is an area of intense scientific focus in the fields of both aging and cancer. In aging, the DNA of telomeres eventually erodes faster than telomerase and its accessory proteins can maintain it, and cells die. In cancer, tumor cells hijack the process, subverting the natural method by which our bodies limit cell growth; cancer cells, then, can grow and multiply unchecked.
One of these accessory proteins, Cdc13, is integral to telomere maintenance and essential for cell viability in yeast, according to researchers at The Wistar Institute. In a study published in the journal Structure, available online now, a team of scientists led by Emmanuel Skordalakes, Ph.D., an associate professor in The Wistar Institute Cancer Center's Gene Expression and Regulation Program, has determined how mutations in a particular region of Cdc13 can lead to defects in telomeres that could jeopardize DNA.
Cdc13 normally functions as a matched-set, where two copies of the protein form what is known as a dimer. Skordalakes found that mutations in a region of Cdc13 (called OB2) prevent Cdc13 copies from binding to each other. The findings help explain the biology of this key telomere maintenance protein, and may eventually lead to novel anticancer therapeutic if their findings translate to a similar molecular system used to maintain human telomeres, Skordalakes says.
"If we could target the OB2 region of Cdc13, for example, it would throw a wrench in the works of telomere maintenance," said Skordalakes. "If you can disrupt recruitment of telomerase in humans, you could potentially drive cells to death."
Cdc13 serves a dual function in telomere replication. First, it keeps the cells' natural DNA repair mechanisms from mistaking the telomere for a broken stretch of DNA, which could cause genetic mayhem if such repair proteins fuse the ends of two chromosomes together, for example. Secondly, Cdc13 recruits telomerase and related proteins to place in order to lengthen the telomeres.
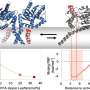
In yeast, telomeres are decorated by a multi-protein complex called CST, which contains the proteins Cdc13 (C), Stn1 (S), and Ten1 (T). Cdc13 is a key member of that complex and serves both to cap the telomere structure and recruit key enzymes.
Skordalakes' newly determined structure demonstrates that, like three of the other four regions of Cdc13, OB2 adopts what is called an oligonucleotide/oligosaccharide-binding fold (OB). These folds normally allow proteins to bind DNA or sugars, but OB2 does neither; its crystal structure indicates that this fold actually forms a large binding surface that helps two Cdc13 proteins to form a dimeric complex.
The authors then used biochemical analyses to determine that OB2 also does not directly bind the protein Stn1. Nevertheless, full-length Cdc13 OB2 mutants deficient in dimerization are also deficient in Stn1 recruitment. When the team inserted strategic Cdc13 mutations into yeast, they found that the cells had abnormally long telomeres, probably as a result of disrupted CST complex assembly caused by impaired Cdc13 dimerization.
"The dimerization of OB2 is required for the proper assembly of the CST complex at the telomeres," Skordalakes says. "When you disrupt oligomerization of this domain, you disrupt assembly of this complex, and thus regulation of telomere length."
Journal information: Structure
Provided by The Wistar Institute