This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New CRISPR tool accelerates and optimizes genome editing
CRISPR/Cas systems have undergone tremendous advancement in the past decade. These precise genome editing tools have applications ranging from transgenic crop development to gene therapy and beyond. And with their recent development of CRISPR-COPIES, researchers at the Center for Advanced Bioenergy and Bioproducts Innovation (CABBI) are further improving CRISPR's versatility and ease of use.
"CRISPR-COPIES is a tool that can quickly identify appropriate chromosomal integration sites for genetic engineering in any organism," said Huimin Zhao, CABBI Conversion Theme Leader and Steven L. Miller Chair of Chemical and Biomolecular Engineering (ChBE) at the University of Illinois.
"It will accelerate our work in the metabolic engineering of non-model yeasts for cost-effective production of chemicals and biofuels."
Gene editing has revolutionized scientists' capabilities in understanding and manipulating genetic information. This form of genetic engineering allows researchers to introduce new traits into an organism, such as resistance to pests or the ability to produce a valuable biochemical.
With CRISPR/Cas systems, researchers can make precise, targeted genetic edits. However, locating optimal integration sites in the genome for these edits has been a critical and largely unsolved problem. Historically, when researchers needed to determine where to target their edits, they would typically manually screen for potential integration sites, then test the site by integrating a reporter gene to assess its cellular fitness and gene expression levels. It's a time- and resource-intensive process.
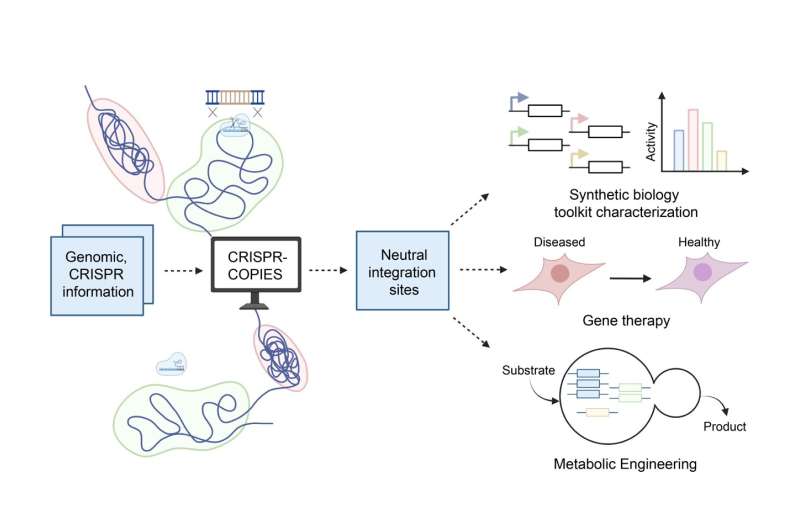
To address this challenge, the CABBI team developed CRISPR-COPIES, a COmputational Pipeline for the Identification of CRISPR/Cas-facilitated intEgration Sites. This tool can identify genome-wide neutral integration sites for most bacterial and fungal genomes within two to three minutes.
"Finding the integration site in the genome manually is like searching for a needle in a haystack," said Aashutosh Boob, a ChBE Ph.D. student at the University of Illinois and primary author of the study. "However, with CRISPR-COPIES, we transform the haystack into a searchable space, empowering researchers to efficiently locate all the needles that align with their specific criteria."
In their paper published in Nucleic Acids Research, the researchers demonstrated the versatility and scalability of CRISPR-COPIES by characterizing integration sites in three diverse species: Cupriavidus necator, Saccharomyces cerevisiae, and HEK 293T cells. They used integration sites found by CRISPR-COPIES to engineer cells with increased production of 5-aminolevulinic acid, a valuable biochemical that has applications in agriculture and the food industry.
In addition, the team has created a user-friendly web interface for CRISPR-COPIES. This incredibly accessible application can be used by researchers even without significant bioinformatics expertise.
A primary objective of CABBI is the engineering of non-model yeasts to produce chemicals and fuels from plant biomass. Economically producing biofuels and bioproducts from low-cost feedstocks at a large scale is a challenge, however, due to the lack of genetic tools and the cumbersome nature of traditional genome-editing methods.
By enabling researchers to swiftly pinpoint genomic loci for targeted gene integration, CRISPR-COPIES provides a streamlined pipeline that facilitates the identification of stable integration sites across the genome. It also eliminates the manual labor involved in designing components for CRISPR/Cas-mediated DNA integration.
For crop engineering, the tool can be used to increase biomass yields, pest resistance, and/or environmental resilience. For converting biomass to valuable chemicals—for instance, by using the yeast S. cerevisiae—CRISPR-COPIES can be used to engineer cells with significantly greater yields.
This versatile software is designed to simplify and accelerate the strain construction process, saving researchers both time and resources. Researchers around the world in both academia and industry can benefit from its utility in strain engineering for biochemical production and transgenic crop development.
More information: Aashutosh Boob et al, CRISPR-COPIES: An in silico platform for discovery of neutral integration sites for CRISPR/Cas-facilitated gene integration, Nucleic Acids Research (2024). DOI: 10.1093/nar/gkae062
Journal information: Nucleic Acids Research
Provided by University of Illinois at Urbana-Champaign