This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
CRISPR/Cas9's role in fine-tuning miRNA expression in tetraploid potatoes
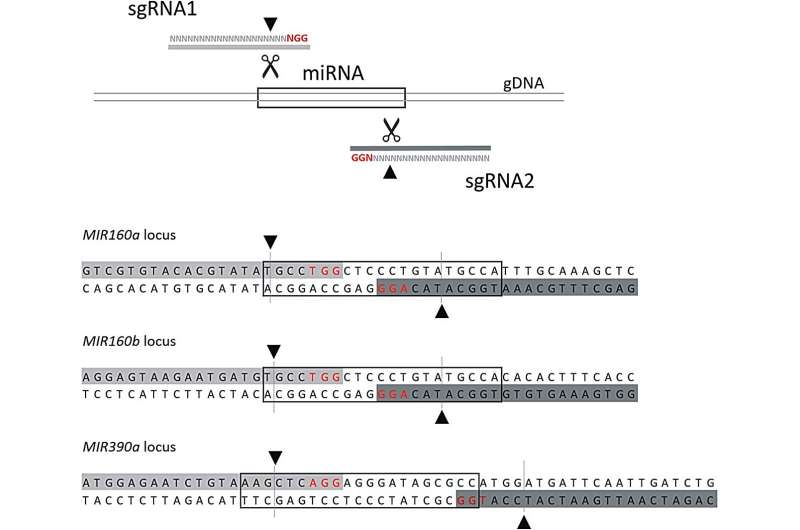
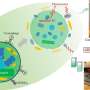
MicroRNAs (miRNAs) play a crucial role in plant gene regulation, with their biogenesis involving complex processes. Fine-tuning miRNAs is a powerful biotechnology strategy that can improve plant performance in the field, such as adjusting crop tolerance to abiotic or biotic stress.
The regulation of RNA by CRISPR/Cas9 mediated gene editing technology is currently a hot research topic. The tetraploid cultivated potato (Solanum tuberosum L.) is one of the most important crops in the world, and CRISPR/Cas9-mediated miRNA editing in potatoes remains unexplored.
In June 2022, Horticulture Research published a research article titled "CRISPR/Cas9-mediated fine-tuning of miRNA expression in tetraploid potato ."
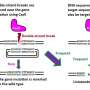
The research utilized a comprehensive CRISPR-Cas9 pipeline to modulate miRNA expression in potatoes, integrating both protoplast transient transfection and Agrobacterium-mediated stable transformation.
The results indicate a correlation between miRNA abundance and mutation types, aligning with pre-miRNA processing efficiency predictions. High-resolution melting analysis proved effective for assessing sgRNA functionality, revealing diverse mutations in transfected protoplasts.
Transfected protoplasts were analyzed to identify potential CRISPR mediated mutations. The results confirm the presence of mutations in MIR160a and MIR160b, while the researchers did not detect any mutations in MIR390a.
In stably transformed potato plants, diverse mutations were observed in different genotypes and transgenic lines, with deletions and insertions being the most common. Notably, mutations were not always at the predicted cutting sites, and some transgenic lines showed all alleles mutated, while others retained non-mutated alleles.
The transformation efficiency and mutation rates varied between potato cultivars and Agrobacterium strains used. The study also found that miRNA abundance correlated with mutation types and numbers. Despite mutations, some miRNA expression persisted, likely due to certain mutations not disrupting pre-miRNA processing.
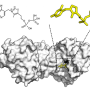
Furthermore, the research introduced new miRNA variants through CRISPR-Cas9 induced mutations. Sequencing revealed these novel variants, with some retaining functionality. However, target analysis showed that most variants had altered target binding, impacting their biological function. Reduced mature miRNA levels due to mutations correlated with negative regulation of their targets.
Finally, the high editing efficiency was confirmed by detecting transgenes, Cas9 expression, and mutations in all transgenic lines.
In conclusion, this study demonstrates the effective use of CRISPR/Cas9 for fine-tuning miRNA expression in polyploid species like potatoes, surpassing the limitations of traditional knock-out mutants. This method's efficiency was validated across different potato genotypes and offers significant potential for precision breeding and plant physiology modulation.
More information: Tjaša Lukan et al, CRISPR/Cas9-mediated fine-tuning of miRNA expression in tetraploid potato, Horticulture Research (2022). DOI: 10.1093/hr/uhac147
Journal information: Horticulture Research
Provided by Plant Phenomics