This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Investigating the role of loops, flags and DNA tension in the spatial organization of chromosomes
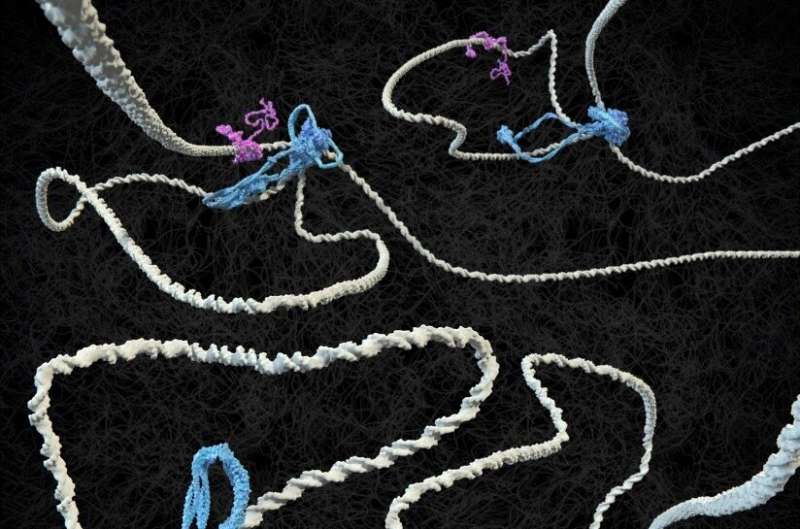
Two protein complexes carry the major responsibility for the spatial organization of chromosomes in our cell nuclei, and DNA tension plays a surprising role in this.
Nanoscientist Cees Dekker and his Ph.D. candidate Roman Barth of the Kavli Institute of Nanoscience at TU Delft, together with Austrian colleagues, have published this finding in the journal Nature on April 19.
Cohesin loops DNA
It has been known for more than a century that the long DNA strands in cell nuclei are neatly folded into the characteristic shape of chromosomes, which resemble bottlebrushes, in preparation for cell division. And between divisions, chromosomes are organized into loops that are important for regulating the processing of genetic information.
In 2018, Dekker and his group were the first to visualize how SMC protein complexes such as condensin and cohesin extrude loops in DNA.
CTCF flags have a direction and determine where a loop begins and ends
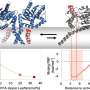
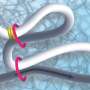
The DNA-binding protein CTCF was found to play a key role in the positioning of loops along the genome. Dekker states, "If you think of DNA as a rope, onto which CTCF flags are pinned at two points, cohesin makes the loops from one flag to the other, but only if the CTCF is oriented correctly. Only one side of the CTCF protein is able to interact with cohesin. Then again, it doesn't always do this, because we thought CTCF would also fail frequently. But now we have measured it. The interaction between the two proteins turns out to be much more subtle than we predicted."
That CTCF and cohesin work together to establish loop boundaries has become basic knowledge in the field, says Ph.D. candidate Roman Barth. "In every conference presentation I attended in the past year, the basic premise was that the cohesin complex extrudes loops between correctly oriented CTCF molecules. But nobody had ever seen in detail how that happens. We have now been able to visualize the essence of this."
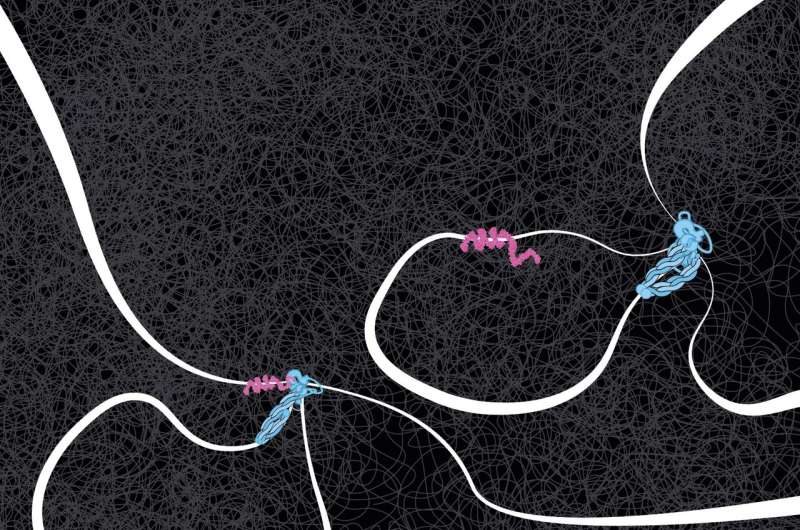
DNA tension plays a surprising role
Colleagues in Jan-Michael Peters' group at the Institute of Molecular Pathology in Vienna succeeded in making the proteins available in pure form. The two ends of a DNA molecule were attached to a surface; the DNA and proteins were stained with a fluorescent dye. The researchers then made an unusual discovery.
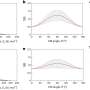
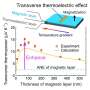
"In the data, Roman discovered that it made a difference whether the DNA strand was very loose or under tension. Without tension, cohesin often ignored the CTCF flag, even if correctly oriented, but when the DNA was under more tension, the CTCF acted as a perfect barrier. So, under the influence of DNA tension, CTCF becomes like a smart traffic light, allowing cohesin to pass or not, depending on the local traffic situation," said Dekker.
When cohesin collides with a CTCF protein, it can stop or continue. The researchers saw that it can also turn around, or even dissolve altogether. How and why this happens are the next questions Dekker and the team hope to answer.
More information: Jan-Michael Peters, CTCF is a DNA-tension-dependent barrier to cohesin-mediated loop extrusion, Nature (2023). DOI: 10.1038/s41586-023-05961-5. www.nature.com/articles/s41586-023-05961-5
Journal information: Nature
Provided by Delft University of Technology