Looping the genome—how cohesin does the trick
Twenty years ago, the protein complex cohesin was first described by researchers at the IMP. They found that its shape strikingly corresponds to its function: when a cell divides, the ring-shaped structure of cohesin keeps sister-chromatids tied together until they are ready to separate.
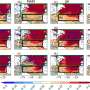
Apart from this important role during cell-divison, other crucial functions of cohesin have been discovered since - at the IMP and elsewhere. One of them is to help fold the DNA, which amounts to about two meters per nucleus, into a compact size by way of creating loops. "We think that the cohesin-ring clamps onto the DNA-strand to hold the loops in place", says IMP-director Jan-Michael Peters whose team worked on the project.
The chromatin-loops are not folded at random. Their exact shape and position play an important role in gene regulation, as they bring otherwise distant areas into close contact. "For a long time, scientists were mystified by how regulatory elements – the enhancers – are able to activate distant genes. Now we think we know the trick: precisely folded loops allow enhancers to come very close to the genes they need to regulate", says Peters. Research results point to cohesin as mediator of this process. Jan-Michael Peters and his team have already shown that the cohesin complex accumulates in areas where loops are formed.
Several scientists recently proposed a so-called "loop-extrusion mechanism" for the folding of chromatin. According to this hypothesis, cohesin is loaded onto DNA at a random site. The DNA strain is then fed through the ring-shaped complex until it encounters a molecular barrier. This element, a DNA-binding protein named CTCF, acts much like a knot tied in a rope and stops the extrusion-process at the correct position. Defined genome-sequences that were previously located far apart are now next to each other and can interact to regulate gene expression.
In Nature online this week, IMP-researchers publish data that support the existence of such a mechanism. First author Georg Busslinger, a PhD-student in Jan-Michael Peters' team, showed in mouse cells that cohesin is indeed translocated on DNA over long distances and that the movement depends on transcription, suggesting that this may serve as a 'motor'.
"The loop extrusion hypothesis has opened up a whole new research area in cell biology and we will probably see many more papers published on this topic in the future", comments Jan-Michael Peters. Understanding cohesin-function is also relevant from a medical perspective since a number of disorders, including certain cancers, are associated with malfunctions of the protein-complex.
More information: Georg A. Busslinger et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl, Nature (2017). DOI: 10.1038/nature22063
Journal information: Nature
Provided by Research Institute of Molecular Pathology