This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Using a genetic algorithm to uncover new catalyst molecules for organic synthesis
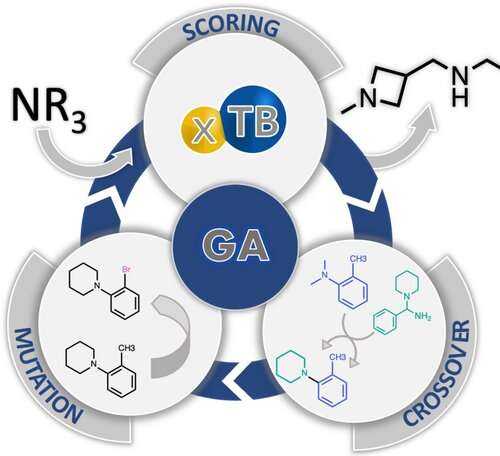
A team of researchers have used a computational method inspired by evolution to discover an organic catalyst with better performance than known catalysts. As the team reports in the journal Angewandte Chemie International Edition, a genetic algorithm suggested new, catalytically active molecular structures for a popular reaction in organic synthesis. The method could be applied more broadly to the search for better molecular catalysts, the team says.
Machine learning systems can already predict material properties and molecular structures in a variety of fields of chemistry with high precision. However, automating the search for new and improved catalysts has to date not been possible, although developing new catalysts for chemical reactions is currently one of the most important goals in chemical research. More efficient catalysts open the door to quicker and easier reactions that consume less energy and form fewer by-products.
The reason behind the difficulties encountered by automated systems when seeking new catalysts lies in reaction transition states, as Jan Halborg Jensen, Professor of computational chemistry at the University of Copenhagen, Denmark, and corresponding author for the study, explains. This is because catalysts influence the transition state; in other words, the moment in a reaction that decides whether a product will be formed or not. The fleeting nature of this moment, and the complexity of the structures formed, with many molecules interacting at the same time, make it difficult to develop models.
To overcome this, Jensen and the team turned to a selection method based on the principles of evolution. A genetic algorithm was used to evaluate a set of starting molecules for fitness for catalyzing the Morita–Baylis–Hillman (MBH) reaction. "Then you take the fittest molecules and mate them, which means that you cut the two parents in random places and recombine fragments from each parent," Jensen explains. "If you do this enough times, the final population can look very different from the initial population, much like a chihuahua differs from its wolf ancestors."
In this way, the final molecules generated by the computer had a new structural motif, a four-membered azetidine ring, which was not present in the initial population. The team then synthesized one of the computer-evolved azetidine candidates and tested it in the reaction, finding that it performed considerably better than the traditional catalyst, DABCO (1,4-diazabicyclo[2.2.2]octane). "Azetidines had never been considered as catalysts for the MBH reaction, so the algorithm made a genuinely novel discovery," says Jensen, highlighting the importance of computer-assisted discoveries in chemical research.
Jensen says that an essential prerequisite for the use of this technique in the future is the knowledge of the key transition state for the reaction in question. He believes that if this is known, genetic algorithms could help to identify new and improved organocatalysts.
More information: Julius Seumer et al, Computational Evolution Of New Catalysts For The Morita–Baylis–Hillman Reaction**, Angewandte Chemie International Edition (2023). DOI: 10.1002/anie.202218565
The data and code are available at sid.erda.dk/sharelink/hGBkdGdCy7 and github.com/jensengroup/mbh_catalyst_ga respectively.
Journal information: Angewandte Chemie International Edition
Provided by Wiley