This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Novel gene discovery points way to better alkaline tolerance in crops
Chinese scientists have identified a key gene involved in crop alkaline tolerance that may greatly improve crop yield in sodic environments through the use of genetic engineering.
This study, led by Prof. Xie Qi's team from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences, in collaboration with seven other institutions, has been published in Science.
Currently, there are more than one billion hectares of saline and alkaline soil in the world, about 60% of which are classified as highly sodic. As a result, the development of saline- and alkaline-resistant crops is an urgent global challenge. However, alkaline tolerance in plants has not been well studied.
Sorghum originated from the harsh environments of Africa and has evolved greater tolerance to multiple abiotic stresses than other major crops (e.g., wheat, rice and maize, etc.). Like some halophytes, sorghum can even survive in a sodic soil with a pH up to 10.0.
In this work, the researchers first performed a genome-wide association study in a diverse sorghum panel and identified an important locus, Alkaline tolerance 1 (AT1), that encodes an atypical G protein γ subunit and controls alkaline tolerance. The AT1 gene has orthologs in other plants; in rice it was named GS3.
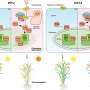
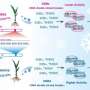
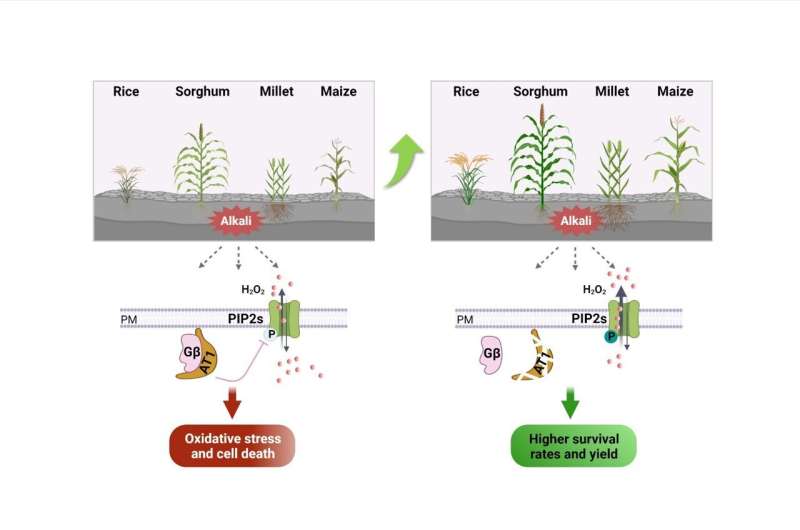
Further experiments confirmed that the at1/gs3 allele produces a C-terminus truncation protein that contributes to the negative alkaline tolerance effect, while knocking out AT1/GS3 (with GS3 being the rice ortholog of AT1) conservatively increased tolerance to alkaline stress in monocot crops, including sorghum, millet, rice and maize.
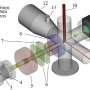
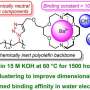
They found that aquaporins PIP2s in reactive oxygen species homeostasis may be involved in Gγ protein signaling. Genetic and cell biological analysis showed that Gγ negatively regulates the phosphorylation of PIP2;1, and the phosphorylation of aquaporins could modulate the efflux of H2O2, leading to decreased ROS levels in plants under alkaline stress.
To evaluate the application of the AT1/GS3 gene in crop production, field trials were carried out in saline and alkaline soils. They found that nonfunctional mutants in a number of monocots, including sorghum, millet, rice and maize, can significantly improve the field performance of crops in terms of biomass or yield production than their not modified controls when grown in sodic soils.
In conclusion, the researchers discovered that an atypical Gγ subunit negatively regulates alkaline stress via modulating the efflux of H2O2 under environmental stress.
"We have discovered the molecular mechanism of a 'star' G-protein that plays a novel role in controlling plant stress response and its downstream molecule aquaporins in H2O2 export," said Prof. Xie.
In addition to illustrating an ecologically important molecular mechanism, this study has great potential for guiding the breeding of alkaline salt-tolerant crops for marginal lands. In this way, it could contribute to global food security as there are more than one billion hectares of saline land worldwide.
More information: Huili Zhang et al, A Gγ protein regulates alkaline sensitivity in crops, Science (2023). DOI: 10.1126/science.ade8416. www.science.org/doi/10.1126/science.ade8416
Journal information: Science
Provided by Chinese Academy of Sciences