This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
SARS-CoV-2 evolved to counter weakness caused by the virus's initial mutation, say researchers
Researchers at Johns Hopkins Medicine say their new studies suggest that the first pandemic-accelerating mutation in the SARS-CoV-2 virus, which causes COVID-19, evolved as a way to correct vulnerabilities caused by the mutation that started the SARS-CoV-2 pandemic.
The new evidence, published in the Dec. 23 issue of Science Advances, addresses important biological questions about two key mutations in the virus's surface "spike" protein, say the researchers. It suggests that a mutation called D614G in the gene for the spike protein, which arose just a few months after the virus started to spread in human populations, was not an adaptation to its new human host.
Rather, the mutation was an adaptation to the dramatic changes that happened in the spike gene just before the start of the pandemic, which allowed SARS-CoV-2 to spread between people by respiratory transmission.
"This study has revealed that the first two genetic alterations in the evolution of the spike protein in SARS-CoV-2 are connected by their function, and this knowledge can improve our understanding of how the spike protein works and how the virus evolves, with important implications for vaccine design and effectiveness of COVID antibodies," says Stephen Gould, Ph.D., professor of biological chemistry at the Johns Hopkins University School of Medicine, whose lab was studying the basic biology of the virus's spike protein when the study began.
The initial mutation in the virus, Gould says, is known by scientists as the furin cleavage site insertion mutation.
Research by other scientists across the world has shown that this mutation enabled the virus's spike protein to be cut and primed it for rapid infection of cells lining the airway.
While this initial mutation was essential in helping SARS-CoV-2 efficiently slip into human cells, the mutation's effects weren't all good, says Gould, as it cut the spike protein structure into two separate pieces.
This change, says Gould, disrupted other functions of the spike protein and created evolutionary pressure for a second mutation that could correct the disrupted functions of the spike protein while maintaining the rapid infection benefits of the initial mutation.
Soon after the pandemic began, in early 2020, researchers from the University of Toronto discovered a subsequent SARS-CoV-2 mutation, called D614G; however, its precise function was not known.
Gould, first author and graduate student Chenxu Guo, and the research team set out to understand the D614G mutation and its effect.
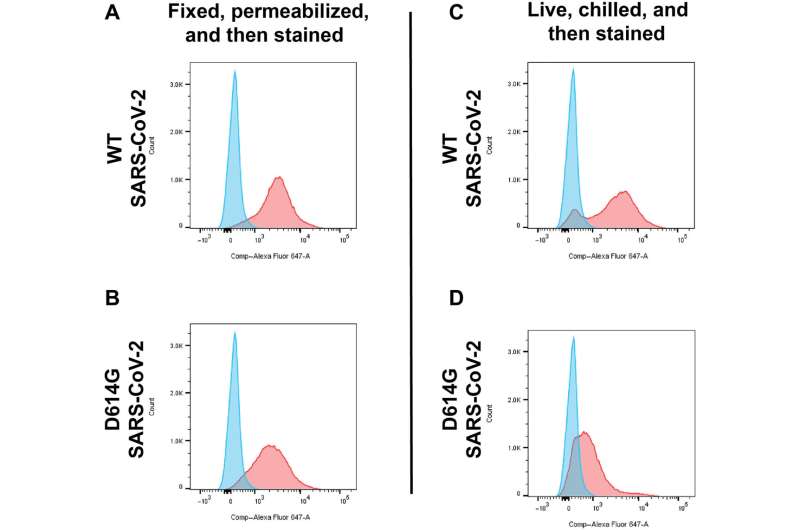
Working with dozens of blood samples from patients with COVID hospitalized in April 2020 at The Johns Hopkins Hospital, Gould's team isolated antibodies for the spike protein from the patients' blood samples. Then, they used these antibodies to track the location of spike proteins in human cells genetically engineered to produce the spiky surface molecules.
They found that the D614G mutation redirects the spike protein and pulls the virus from the surface of human cells into a tiny compartment within the cell called a lysosome, which the spike protein reprograms into storage containers that are used to release infectious virus particles from the cell.
In addition, the D614G mutation caused a three-fold drop in the abundance of spike proteins at the cell surface.
"With less spike protein on the surface of virus-infected cells, it may be more difficult for the immune system to identify and kill those virus-containing cells," says Gould.
Gould also collaborated with the laboratory of Andrew Pekosz, Ph.D., professor of microbiology and immunology at the Johns Hopkins Bloomberg School of Public Health, who confirmed that these mutations have the same effects in cells infected with SARS-CoV-2.
The researchers caution that the study does not provide information about the still-debated origins of the virus. However, their work suggests that the two mutations likely arose in rapid succession.
Gould and Pekosz are continuing their studies by testing how spike protein mutations in more recent virus strains affect spike protein trafficking, studying the identity of the human proteins that deliver spike proteins to lysosomes, and researching how spike proteins convert lysosomes into compartments that release more virus.
More information: Chenxu Guo et al, The D614G mutation redirects SARS-CoV-2 spike to lysosomes and suppresses deleterious traits of the furin cleavage site insertion mutation, Science Advances (2022). DOI: 10.1126/sciadv.ade5085
Journal information: Science Advances
Provided by Johns Hopkins University School of Medicine