How a shape-shifting receptor influences cell growth
Receptors found on cell surfaces bind to hormones, proteins, and other molecules, helping cells respond to their environment. MIT chemists have now discovered how one of these receptors changes its shape when it binds to its target, and how those changes trigger cells to grow and proliferate.
This receptor, known as epidermal growth factor receptor (EGFR), is overexpressed in many types of cancer and is the target of several cancer drugs. These drugs often work well at first, but tumors can become resistant to them. Understanding the mechanism of these receptors better may help researchers design drugs that can evade that resistance, says Gabriela Schlau-Cohen, an associate professor of chemistry at MIT.
"Thinking about more general mechanisms to target EGFR is an exciting new direction, and gives you a new avenue to think about possible therapies that may not evolve resistance as easily," she says.
Schlau-Cohen and Bin Zhang, the Pfizer-Laubach Career Development Assistant Professor of Chemistry, are the senior authors of the study, which appears today in Nature Communications. The paper's lead authors are MIT graduate student Shwetha Srinivasan and former MIT postdoc Raju Regmi.
Shape-changing receptors
The EGF receptor is one of many receptors that help control cell growth. Found on most types of mammalian epithelial cells, which line body surfaces and organs, it can respond to several types of growth factors in addition to EGF. Some types of cancer, especially lung cancer and glioblastoma, overexpress the EGF receptor, which can lead to uncontrolled growth.
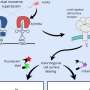
Like most cell receptors, the EGFR spans the cell membrane. An extracellular region of the receptor interacts with its target molecule (also called a ligand); a transmembrane section is embedded within the membrane; and an intracellular section interacts with cellular machinery that controls growth pathways.
The extracellular portion of the receptor has been analyzed in detail, but the transmembrane and intracellular sections have been difficult to study because they are more disordered and can't be crystallized.
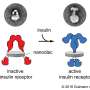
About five years ago, Schlau-Cohen set out to try to learn more about those lesser-known structures. Her team embedded the proteins in a special type of self-assembling membrane called a nanodisc, which mimics the cell membrane. Then, she used single molecule FRET (fluorescence resonance energy transfer) to study how the conformation of the receptor changes when it binds to EGF.
FRET is commonly used to measure tiny distances between two fluorescent molecules. The researchers labeled the nanodisc membrane and the end of the intracellular tail of the protein with two different fluorophores, which allowed them to measure the distance between the protein tail and the cell membrane, under a variety of circumstances.
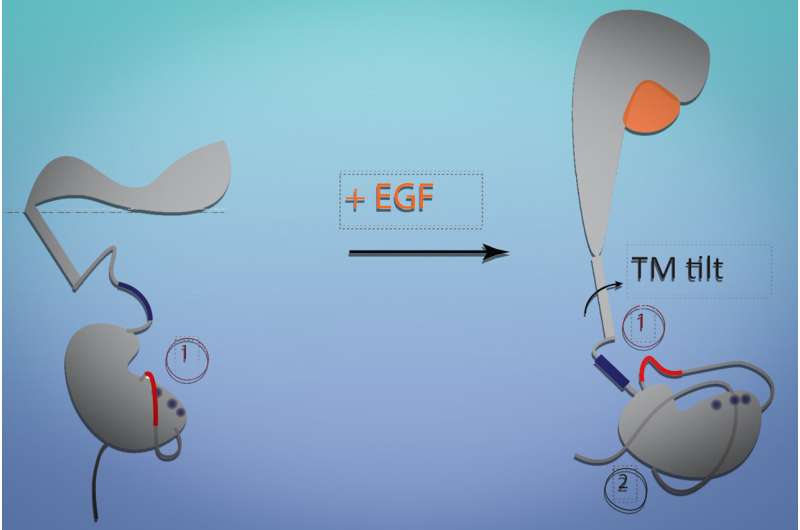
To their surprise, the researchers found that EGF binding led to a major change in the conformation of the receptor. Most models of receptor signaling involve interaction of multiple transmembrane helices to bring about large-scale conformational changes, but the EGF receptor, which has only a single helical segment within the membrane, appears to undergo such a change without interacting with other receptor molecules.
"The idea of a single alpha helix being able to transduce such a large conformational rearrangement was really surprising to us," Schlau-Cohen says.
Molecular modeling
To learn more about how this shape change would affect the receptor's function, Schlau-Cohen's lab teamed up with Zhang, whose lab does computer simulations of molecular interactions. This kind of modeling, known as molecular dynamics, can model how a molecular system changes over time.
The modeling showed that when the receptor binds to EGF, the extracellular segment of the receptor stands up vertically, and when the receptor is not bound, it lies flat against the cell membrane. Similar to a hinge closing, when the receptor falls flat, it tilts the transmembrane segment and pulls the intracellular segment closer to the membrane. This blocks the intracellular region of the protein from being able to interact with the machinery needed to launch cell growth. EGF binding makes those regions more available, helping to activate growth signaling pathways.
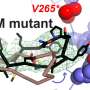
The researchers also used their model to discover that positively charged amino acids in the intracellular segment, near the cell membrane, are key to these interactions. When the researchers mutated those amino acids, switching them from charged to neutral, ligand binding no longer activated the receptor.
"There's a nice consistency we can see between the simulation and experiment," Zhang says. "With the molecular dynamics simulations, we can figure out what are the amino acids that are essential for the coupling, and quantify the role of different amino acids. Then Gabriela showed that those predictions turned out to be correct."
The researchers also found that cetuximab, a drug that binds to the EGF receptor, prevents this conformational change from occurring. Cetuximab has shown some success in treating patients with colorectal or head and neck cancer, but tumors can become resistant to it. Learning more about the mechanism of how EGFR responds to different ligands could help researchers to design drugs that might be less likely to lead to resistance, the researchers say.
More information: Nature Communications (2022). DOI: 10.5281/zenodo.6564353
Journal information: Nature Communications
Provided by Massachusetts Institute of Technology