Understanding the genomic modifications in transgenic papaya

The transgenic papaya "SunUp" was developed in the 1990s and was widely publicized because of its ability to resist the papaya ringspot virus. Although researchers had identified the genomic sequence of SunUp by 2008, it was unclear where the transgenic insertions were and what effect they had. A new study has identified these changes and how they influence the plants.
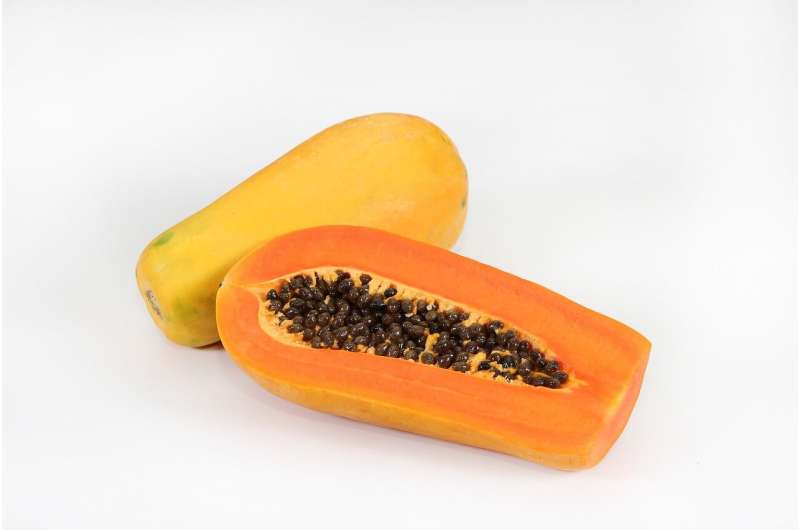
Papaya fruits are a rich source of potassium, magnesium, and vitamins A and C, propelling a steady increase in their global production. Papaya originated and was domesticated in southern Mexico and Central America and is now cultivated in tropical and subtropical regions worldwide.
Wild papaya has small seedy fruits with very little edible flesh, while the domesticated version can weigh more than five pounds. However, there had been one major problem: Papaya was susceptible to the papaya ringspot virus, resulting in stunted plants that did not produce mature fruit, and there was no resistance in the papaya genetic code.
Then researchers developed the transgenic papaya SunUp by using a technique called particle bombardment-mediated transformation. Gold particles were covered with the coat protein gene of the virus and shot into the cells of the non-transgenic papaya "Sunset" using a gene gun. SunUp therefore contained gene sequences of the virus and was protected from infection.
"It took us 8 years to read each DNA nucleotide in the insertions and rearrangements, and we repeated the sequencing using different technologies to understand the nature of these transgenic insertions," said plant biologist Ray Ming of the University of Illinois Urbana-Champaign. "The insertion was so complex that although we sequenced the genome in 2008, we didn't know where the transgenic sequences were located."
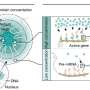
The group discovered that SunUp had an insertion of 1.6 million base pairs, which consisted of DNA fragments not only from the gene gun, but also nuclear genome fragments and organelle genome fragments from the progenitor Sunset cultivar. The results are published in Nature Genetics.
"This research nicely unravels the changes caused by particle bombardment-mediated transformation in the papaya genome and provides new insights into the domestication of papaya," said Gerald Schoenknecht, a program director in NSF's Division of Integrative Organismal Systems.
Although there is such a large insertion, the transgenic manipulation did not cause any change in gene expression, the researchers said. "We looked at every gene sequence and there is no impact on the genome function," Ming said. "When we compared SunUp and Sunset, they only have 20 genes that are differentially expressed, which are due to transposon-mediated rearrangements and not from the genetic manipulation done by particle bombardment-mediated transformation."
Transposon-mediated rearrangements occur naturally and lead to gradual changes over time, which is expected since SunUp and Sunset have been growing and diverging for 30 years.
More information: Jingjing Yue et al, SunUp and Sunset genomes revealed impact of particle bombardment mediated transformation and domestication history in papaya, Nature Genetics (2022). DOI: 10.1038/s41588-022-01068-1
Journal information: Nature Genetics
Provided by National Science Foundation