Mapping how cell types and tissues develop
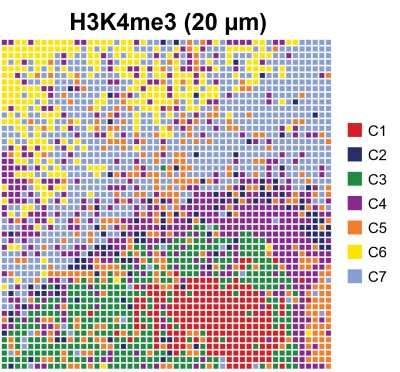
Researchers have developed a technique that allows them to look simultaneously at a spatial level and at a genome-wide level at epigenetic mechanisms underlying tissue development, a breakthrough with multiple scientific and medical applications.
While current technologies allow scientists to examine epigenetic information in the bulk of a tissue, and recent breakthroughs further extend to an individual cell level, it's currently not possible to visualize them directly in a tissue with cellular resolution. This is a critical obstacle, since there's a strong connection between how cells are organized in tissue and how they function. In a new study published in Science, a team of researchers have developed a technique to enable for the first time the spatial mapping of epigenetic regulations connected to development and disease. Led by Rong Fan, professor of biomedical engineering at Yale University and of pathology at Yale School of Medicine, the study was done in collaboration with researchers at Karolinska Institutet in Stockholm, Sweden.
We all start with the same genetic material in a single cell, but that cell quickly multiplies into trillions of different cells. Each cell has the same genome, but the difference lies in which genes have been activated or repressed. Different chemical epigenetic modifications in the embryo steer the development—switching certain genes on and other ones off—and allow for the formation of different organs. Epigenetics is what regulates the expression of those genes. For their study, the researchers focused on one of the most significant epigenetic changes, known as histone modifications.
"What underlying mechanisms control the cell types and tissue development?" Fan said. "That's what we're able to 'see' now with this new technology—spatial-CUT&Tag. "This is the first time we can look at which histone modifications control genome-wide gene expression directly in the tissue, and how they do it—it adds a whole new dimension to this emerging field of spatial biology."
Other technologies allow for the profiling of epigenetic modifications in individual cells, but not within their tissue. Being able to track them in their full context provides invaluable insight about cell organization, tissue development and disease. Co-author Gonçalo Castelo-Branco, professor of glial cell biology at Karolinska Institutet, noted that the technology provides information about which genes in a tissue section are switched on and off.
"That gives a lot of additional information—for instance, why is a nerve cell a nerve cell?" he said. "Before, you could check cell by cell, but then you would lose all the information of where the cells were and how they might have been interacting with each other. With this technology, we have now also the spatial location. Moreover, you can determine in that location all this epigenomic information, which is one of the really amazing aspects of the technology."
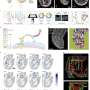
Specifically, the researchers looked at modifications to chromatin, the material that enwraps the DNA of every cell and controls access to it, in the embryonic and postnatal brains of mice. It had been shown that specific histone modifications can control different cortical layers of the brain in the embryonic stage, but it was unclear if they would have the same function after birth.
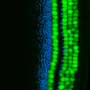
"So now this is the first time we can see those epigenetic features directly in a postnatal brain," Fan said. "We can say, "Wow, actually those epigenetic features are organized in stripes, which correspond to the cortical layers of the mouse brain." That means they might be controlling or maintaining the formation of these layers."
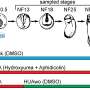
The researchers created their technique by designing microfluidic devices with multiple microchannels that are used to deliver DNA barcodes to the genome DNA where the epigenetic modifications are present. The study's lead author, Yanxiang Deng, a postdoctoral associate in Rong Fan's lab, said the researchers can then retrieve the genome DNA and reconstruct the spatial location by computational methods. If there is a histone modification on a genome DNA, a tagged DNA barcode is associated with that histone modification, allowing the researchers to locate in the tissue the epigenetic information.
The spatial-CUT&Tag technique adds an extra dimension to the CUT&Tag technique developed in recent years, allowing researchers to get extremely precise information, pixel by pixel. The technology enables researchers to place an identifier—like a GPS coordinate—on each cell in the tissue section.
"With that, you can reconstruct the map of the tissue," Castelo-Branco said. "It is the ability to put a coordinate for every single pixel in that tissue section and then you can go back and say "OK, this is where this histone modification is present, in this specific gene, in this specific cell or group of cells.'"
In the long run, the research team hopes that the technology will be used to better understand the development of disease.
"Epigenetic drugs are something that's emerging now, so potentially we can develop drugs to target those epigenetic mechanisms," Fan said. "Having the tools to understand the epigenetic origin of different disease states could open up a whole new avenue of therapeutics."
The researchers emphasized that the technology has a broad range of applications.
"This is a starting point, and it will allow a lot of other findings similar to this on different tissues," Castelo-Branco said.
More information: Yanxiang Deng et al, Spatial-CUT&Tag: Spatially resolved chromatin modification profiling at the cellular level, Science (2022). DOI: 10.1126/science.abg7216
Journal information: Science
Provided by Yale University