Technique allows mapping of epigenetic information in single cells at scale

Histones are tiny proteins that bind to DNA and hold information that can help turn on or off individual genes. Researchers at Karolinska Institutet have developed a technique that makes it possible to examine how different versions of histones bind to the genome in tens of thousands of individual cells at the same time. The technique was applied to the mouse brain and can be used to study epigenetics at a single-cell level in other complex tissues. The study is published in the journal Nature Biotechnology.
"This technique will be an important tool for examining what makes cells different from each other at the epigenetic level," says Marek Bartosovic, post-doctoral fellow at the Department of Medical Biochemistry and Biophysics, Karolinska Institute. "We anticipate that it will be widely implemented by the broad biomedical community in a wide variety of research fields."
Even though all cells in our bodies contain exactly the same genetic information written in DNA, they read and execute this information differently. For example, a neuron in the brain reads the DNA differently than a skin cell or a fat cell. Epigenetics play a crucial role in the interpretation of the genetic information and allows the cells to execute specialized functions.
Modification of histones is one type of epigenetic information. Histones are attached to DNA like beads on a string and decorate it as 'epigenetic stickers.' These stickers label which genes should be turned on or off. All together this system coordinates to ensure that thousands of genes are switched on or off at the right place and time and in the right cell.
To better understand how cells become different and specialized, it is important to know which parts of the DNA are marked by which histone 'stickers' in each individual organ and in each individual cell within that organ. Until recently, it was not possible to look at histone modifications of an individual cell. To examine the histone modifications in one specific cell type, a very high number of cells and cumbersome methods of cell isolation would be required. The final epigenetic histone profile of one cell type would then be an averaged view of thousands of cells.
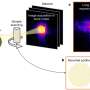
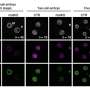
In this study, the researchers describe a method that introduces a new way of looking at histone modifications in unprecedented detail in a single cell and at a large scale. By coupling and further optimizing other epigenomic techniques, the researchers were able to identify gene regulatory principles for brain cells in mice based on their histone profile.
"Our method—single-cell CUT&Tag—makes it possible to examine tens of thousands of single cells at the same time, giving an unbiased view of the epigenetic information in complex tissues with unparalleled resolution," Gonçalo Castelo-Branco, associate professor at the Department of Medical Biochemistry and Biophysics, Karolinska Institute, says. "Next, we would like to apply single-cell CUT&Tag in the human brain, both in development and in various diseases. For instance, we would like to investigate which epigenetic processes contribute to neurodegeneration during multiple sclerosis and whether we would be able to manipulate these processes in order to alleviate the disease."
More information: Marek Bartosovic et al. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nature Biotechnology, online April 12, 2021, DOI: 10.1038/s41587-021-00869-9
Journal information: Nature Biotechnology
Provided by Karolinska Institutet