Scientists determine the mode of action of essential proteins involved in cancer and Alzheimer's disease
The proteins that belong to the HAT family are essential for life as they transport amino acids across the cell membrane. Although the members of this family are practically identical, some transport certain amino acids and not others. This specialization determines their involvement in specific functions, such as cell growth or neuronal functions, and consequently in related diseases like cancer or neurodegenerative conditions such as Alzheimer´s disease. What confers this specificity and diversity of functions? This is one of the questions asked by researchers at the Spanish National Cancer Research Center (CNIO) and the Institute for Research in Biomedicine (IRB Barcelona), who led the study, and one whose answer has been published this week in the journal Proceedings of the National Academy of Sciences (PNAS).
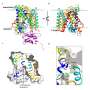
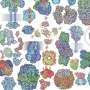
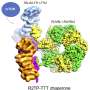
Thanks to the latest high-resolution structural technologies such as cryo-electron microscopy, combined with computational modeling and the design of mutants of these proteins, the researchers have been able to observe the structure of one of the members of this protein family in atomic detail and decipher its function. The results of the study show how only a few residues—located in defined regions—of this family of proteins select the specific amino acids to which they will bind and are therefore responsible for the protein to participate in specific physiological functions.
Armed with this information, the researchers now face the challenge of finding new therapies and diagnostic tools for diseases that involve the HAT family of transporter proteins, with a particular interest in those conditions that pose serious health problems, such as cancer and neurodegenerative disorders such as Alzheimer´s disease.
Understanding how to disrupt their function
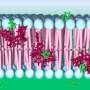
Amino acids, the basic building blocks of life, enter and leave cells, enabling them to grow, divide and develop their functions. This movement into and out of the cell occurs thanks to gates embedded in the cell membrane that are formed by proteins of the HAT family, among others.
Although HAT proteins are practically identical in structure, some transport certain amino acids and not others, thereby conferring each member of the family specific functions, such as the participation in cell growth; a role in diseases such as cancer; neuronal functions; and the transport of toxic substances and involvement in addiction to substances such as cocaine.
To understand this specificity of function, the scientists set out to study the 3D structure of this important family of proteins. "Classical techniques used to determine the structure of proteins, such as those using X-rays, have had limited success with proteins that are embedded in biological membranes, and so many questions have remained unresolved," says Oscar Llorca, head of the Macromolecular Complexes in DNA Damage Response Group at CNIO, Director of the Structural Biology Programme of this Centre and co-author of the work."The combination of structural resolution by cryo-electron microscopy with molecular dynamics calculations and functional studies provides an experimental platform with a lot of potential that allows us to unravel the function of amino acid transporters. In this case, we have applied this technology to identify the molecular mechanisms that lead these proteins to transport some amino acids but not others," says Manuel Palacín, head of the Amino Acid and Disease Transporters laboratory at IRB Barcelona, professor of the University of Barcelona and Unit leader of CIBERER.
New drugs against cancer and Alzheimer´s disease
Thanks to cryo-electron microscopy, a field in which Llorca is an international expert, visualization of the molecular structure of proteins has taken a giant step towards what we now know as the golden age of 3D structures. This new technology, which won the Nobel Prize in Chemistry in 2017, has not only served to observe biological processes like never before but it is also contributing to accelerating the development of new compounds and drugs of interest to treat cancer and other human diseases.
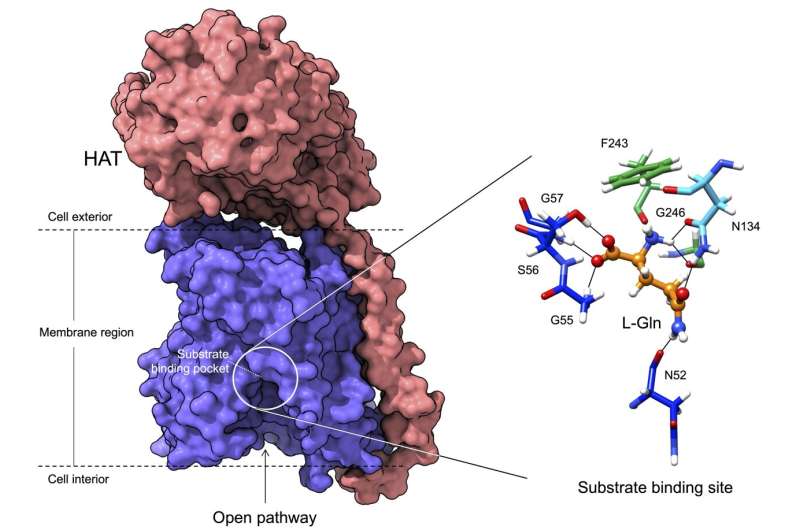
In this work, using cryo-electron microscopy, the researchers have been able to visualize the structure of a member of the HAT family at atomic resolution and determine the pocket where these proteins bind to amino acids, as well as the details of the mechanism by which this recognition occurs.
The atomic details reveal that only a few residues of these proteins determine the amino acids to which they bind and therefore their specific functions. In addition, the study demonstrates how the substitutions of some residues for others in these positions in the different members of the family are responsible for modifying the specificity of recognition and transport of some amino acids and not others.
The results of this research could lead to compounds that act on specific regions of these proteins, and to manage the disorders in which they participate, such as cancer and neurodegenerative diseases like Alzheimer's disease.
More information: Structural basis for substrate specificity of heteromeric transporters of neutral amino acids, Proceedings of the National Academy of Sciences (2021). DOI: 10.1073/pnas.2113573118.
Journal information: Proceedings of the National Academy of Sciences