Scientists discover 'bulkheads' between liver cells
Researchers from Skoltech and their colleagues from Germany and the U.S. have discovered structures responsible for the shape of bile canaliculi, a network of canals in the liver. Published in the Journal of Cell Biology, their study also reveals the gene essential for the formation of these structures, which the team compares to boat bulkheads.
The body incorporates numerous tubular structures, such as blood vessels or the intestine. They are usually formed by many epithelial cells sharing their apical, or "inside," surfaces. By contrast, the most abundant liver cells, called hepatocytes, form lumina by sharing surfaces only with their directly adjacent neighbors. This results in very narrow canals that branch out in a 3D network.
It has so far remained unknown what causes hepatocytes to behave in this way, unlike the other kind of liver cells, cholangiocytes. Those build much larger tubes with a fairly straightforward structure—more like conventional epithelial cells. For a while, some kind of a local mechanical feature or interaction was suspected to account for the shape of the hepatocyte lumina.
Apical bulkheads
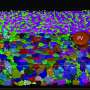
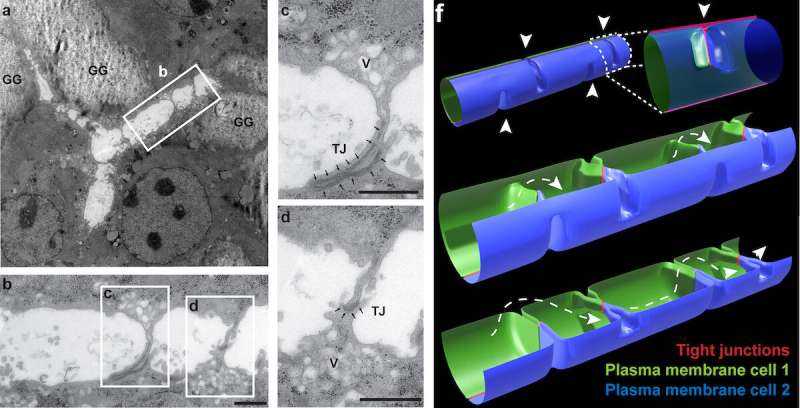
Marino Zerial and his team from the Max Planck Institute of Molecular Cell Biology and Genetics, in collaboration with Skoltech researchers led by Associate Professor Timofei Zatsepin, and their colleagues, have discovered extensions on the apical surfaces of hepatocytes that form structures reminiscent of boat bulkheads in the lumina. The team observed these "reinforcement ribs" with an electron microscope and showed them to be responsible for the narrow shape of bile canaliculi and the intricacy of their networks.
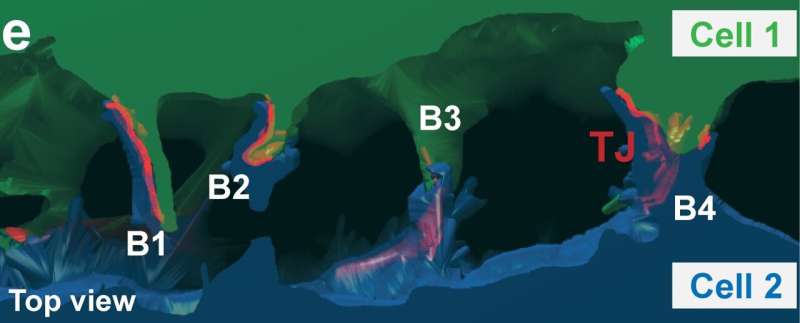
Unlike in a boat hull, the apical extensions do not separate bile canaliculi but preserve their continuity. Yet both structures—the human-made and the cell-intrinsic one—are there to provide rigidity.
To rule out the possibility of apical bulkheads being an artifact of observations in vitro, the team examined an embryonic mouse liver by electron microscopy, confirming the presence of bulkhead-like connections in vivo in the nascent bile canaliculi. The bulkheads did not divide the lumen into isolated chambers. An adult mouse liver also exhibited such structures. In vivo observations allowed the researchers to make sure the newly discovered bulkheads were not in fact confused with microvilli, a common type of cellular membrane protrusion.
Underlying genetics
The team then went on to examine gene candidates potentially responsible for bulkhead formation, eventually focusing the attention on Rab35. The functions of the well-known Rab35 protein were not previously connected to hepatocyte lumen morphology. Electron microscopy and 3D modeling revealed that Rab35 knockdown led to hepatocytes not developing bulkhead-like structures and consequently forming lumina like those of cholangiocytes.
"Hepatocytes and cholangiocytes have one progenitor—the hepatoblast, so the partial phenotype switch of hepatocytes with Rab35 knockdown clearly shows that Rab35 is involved in lumina tuning. But we know that Rab35 does not directly control bulkhead formation. Being a well-known transporter, it moves proteins within the cell," Timofei Zatsepin said. "Rab35 is clearly in charge of transporting some protein complex or complexes that actually drive bulkhead development. We want to uncover the whole precise mechanism that makes the liver so different from other organs."
The researchers suggest that the bulkhead structures they identified can be manipulated for further research with potential medical applications. "These structures are amazing and beautiful by themselves. The quasi-periodic pattern of apical bulkheads, whose distance is in the range of the lumen diameter, looks like the load-bearing structures in human-made constructions," Zatsepin commented, adding that the team is "going to study the contribution of this mechanism to the functioning of the liver and its regeneration in the context of fatty liver disease and fibrosis."
More information: Lenka Belicova et al, Anisotropic expansion of hepatocyte lumina enforced by apical bulkheads, Journal of Cell Biology (2021). DOI: 10.1083/jcb.202103003
Journal information: Journal of Cell Biology
Provided by Skolkovo Institute of Science and Technology