Machine learning plus insights from genetic research shows the workings of cells
We combined a machine learning algorithm with knowledge gleaned from hundreds of biological experiments to develop a technique that allows biomedical researchers to figure out the functions of the proteins that turn genes on and off in cells, called transcription factors. This knowledge could make it easier to develop drugs for a wide range of diseases.
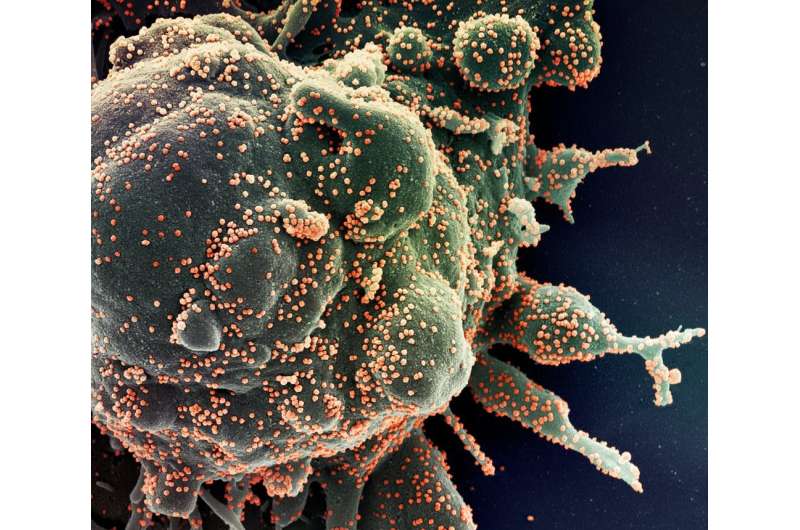
Early on during the COVID-19 pandemic, scientists who worked out the genetic code of the RNA molecules of cells in the lungs and intestines found that only a small group of cells in these organs were most vulnerable to being infected by the SARS-CoV-2 virus. That allowed researchers to focus on blocking the virus's ability to enter these cells. Our technique could make it easier for researchers to find this kind of information.
The biological knowledge we work with comes from this kind of RNA sequencing, which gives researchers a snapshot of the hundreds of thousands of RNA molecules in a cell as they are being translated into proteins. A widely praised machine learning tool, the Seurat analysis platform, has helped researchers all across the world discover new cell populations in healthy and diseased organs. This machine learning tool processes data from single-cell RNA sequencing without any information ahead of time about how these genes function and relate to each other.
Our technique takes a different approach by adding knowledge about certain genes and cell types to find clues about the distinct roles of cells. There has been more than a decade of research identifying all the potential targets of transcription factors.
Armed with this knowledge, we used a mathematical approach called Bayesian inference. In this technique, prior knowledge is converted into probabilities that can be calculated on a computer. In our case it's the probability of a gene being regulated by a given transcription factor. We then used a machine learning algorithm to figure out the function of the transcription factors in each one of the thousands of cells we analyzed.
We published our technique, called Bayesian Inference Transcription Factor Activity Model, in the journal Genome Research and also made the software freely available so that other researchers can test and use it.
Our approach works across a broad range of cell types and organs and could be used to develop treatments for diseases like COVID-19 or Alzheimer's. Drugs for these difficult-to-treat diseases work best if they target cells that cause the disease and avoid collateral damage to other cells. Our technique makes it easier for researchers to home in on these targets.
Single-cell RNA-sequencing has revealed how each organ can have 10, 20 or even more subtypes of specialized cells, each with distinct functions. A very exciting new development is the emergence of spatial transcriptomics, in which RNA sequencing is performed in a spatial grid that allows researchers to study the RNA of cells at specific locations in an organ.
A recent paper used a Bayesian statistics approach similar to ours to figure out distinct roles of cells while taking into account their proximity to one another. Another research group combined spatial data with single-cell RNA-sequencing data and studied the distinct functions of neighboring cells.
We plan to work with colleagues to use our new technique to study complex diseases such as Alzheimer's disease and COVID-19, work that could lead to new drugs for these diseases. We also want to work with colleagues to better understand the complexity of interactions among cells.
Journal information: Genome Research
Provided by The Conversation
This article is republished from The Conversation under a Creative Commons license. Read the original article.![]()