Evolution proteomics approach opens view into how new gene functions arise
The creation of genes with new functions is a major driver of developmental innovation in all living organisms. How these genes acquire new functions over evolutionary time scales, however, is unclear.
Whole genome duplications occur often, giving organisms redundant copies of genes that can mutate and acquire new functionality. These duplicate genes are similar at the sequence level, and it's commonly assumed that as species diverge, these genes maintain the same functions over millions of years. This assumption leads scientists to believe that genes with similar sequences have the same functions, but that may not be true.
Purdue University scientist Dan Szymanski and graduate student Youngwoo Lee have developed a new high-throughput method to analyze these genes and the proteins they encode, identifying functional differences across a range of plant species even among genes that look to be the same. Their work suggests that these otherwise duplicate genes can give rise to new protein functions, as well as new interactions among protein complexes, that drive biological evolution and innovation in plants.
"Most analyses of plant evolution are based on DNA and protein sequences, but our analysis is based on unique functional interactions or protein-protein interactions among related proteins. This goes far beyond sequence and provides deeper functional clues," said Szymanski, a professor in the Department of Botany and Plant Pathology whose findings were published in the journal Science Advances. "We can develop hypotheses about how particular protein-protein interactions might have evolved during a changing environment or as a result of a developmental change in the organism."
Szymanski and Lee's method involves comparing the proteins and protein complexes from multiple plants through mass spectrometry. Using the model plant Arabidopsis thaliana as well as cotton, soybeans and rice—which all share a common ancestor ‑ the scientists detected mass differences in evolutionarily related proteins. That suggests these proteins, which should otherwise be the same in all the different plants, found ways to form new protein complexes and develop new functions. The same family of proteins could then be analyzed across a wide variety of species to test for evolutionary patterns in the protein-protein interaction data.
"As plants evolve and acquire duplications to their genomes, some proteins mutate to develop a function not present in the ancestral gene. We can see that based on distinct masses of protein complexes," Szymanski said. "They bind to other proteins or themselves, and sometimes these differences generate important new functions that are retained widely in the lineage."
While it could be argued these protein-protein interactions formed through random chance, Szymanski's team provides evidence that these developments were driven by environmental circumstances and retained in plants for millions of years.
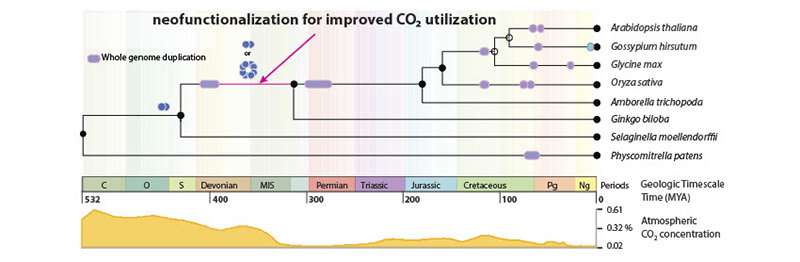
The scientists give the example of carbonic anhydrase, a protein that is key for carbon dioxide transport. This protein would not have limited plant productivity in high-carbon environments. About 400 million years ago, however, carbon dioxide levels in Earth's atmosphere were falling due to the widespread colonization by plants. This new CO2-limiting environment may have made carbonic anhydrase more important, as its neofunctionalization into a more efficient form was traced to this interval in Earth's history.
The process Szymanski and Lee developed provides a molecular explanation of a common path to protein neofunctionalization.
"This reveals which proteins have changed and how protein-protein interactions have evolved," Szymanski said. "That can tell us a lot about the types of proteins that innovated in response to changes in the environment or developmental programs of the plant."
More information: Youngwoo Lee et al. Multimerization variants as potential drivers of neofunctionalization, Science Advances (2021). DOI: 10.1126/sciadv.abf0984
Journal information: Science Advances
Provided by Purdue University


















