Looking through MudPIT for protein interactions

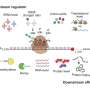
The instructions of life encoded in our genes are decoded through the translation into mRNA, which then instructs the synthesis of proteins. Because mRNA translation is an essential process, it is carefully coordinated through "translational control." However, defects in translational control and protein synthesis lead to many pathologies, which makes mRNA translation an important therapeutic target for many human diseases.
Now, using a technique called Multidimensional Protein Identification Technology (MudPIT), Andrew Link, Ph.D., and colleagues have discovered unexpected protein interactions and sites of a protein modification called phosphorylation involved in mRNA translation.
In a study published in the journal Proteomics, they validated previously described protein interactions involved in mRNA translation, in the model organism yeast (S. cerevisiae).
Their study further identified novel protein interactions and phosphorylation sites for S. cerevisiae's mRNA translation proteins and protein complexes.
These previously unpublished protein interactions and phosphorylation sites can drive future functional, mechanistic, and structural studies towards a better understanding of mRNA translation.
More information: Andrew J. Link et al. Targeted identification of protein interactions in eukaryotic mRNA translation, PROTEOMICS (2020). DOI: 10.1002/pmic.201900177
Journal information: Proteomics
Provided by Vanderbilt University